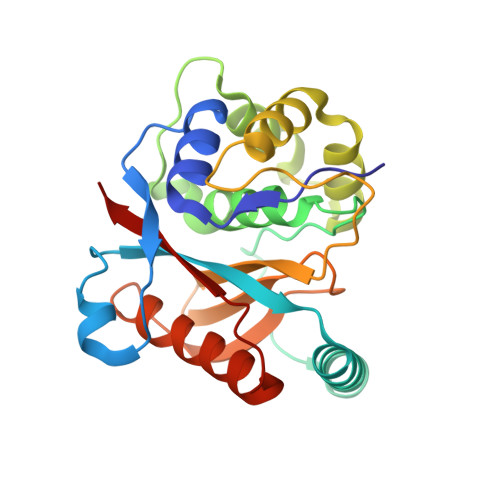
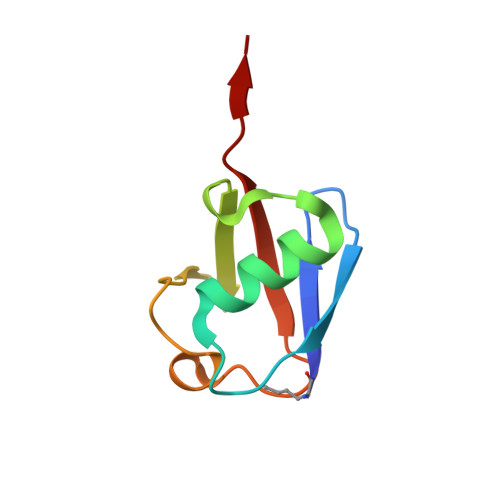
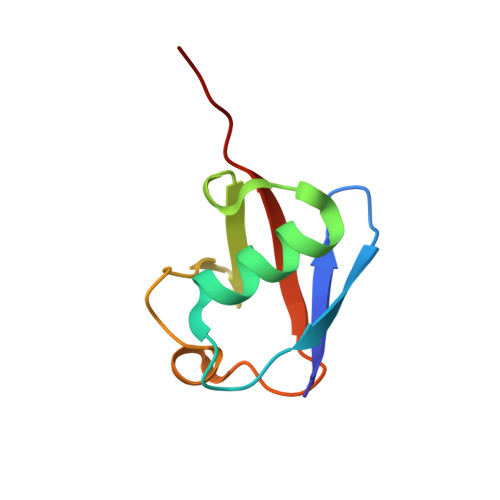
K27-Linked Diubiquitin Inhibits UCHL3 via an Unusual Kinetic Trap.
van Tilburg, G.B.A., Murachelli, A.G., Fish, A., van der Heden van Noort, G.J., Ovaa, H., Sixma, T.K.(2021) Cell Chem Biol 28: 191-201.e8
- PubMed: 33238157 Search on PubMed
- DOI: https://doi.org/10.1016/j.chembiol.2020.11.005
- Primary Citation Related Structures:
6QML - PubMed Abstract:
Functional analysis of lysine 27-linked ubiquitin chains ( K27 Ub) is difficult due to the inability to make them through enzymatic methods and due to a lack of model tools and substrates. Here we generate a series of ubiquitin (Ub) tools to study how the deubiquitinase UCHL3 responds to K27 Ub chains in comparison to lysine 63-linked chains and mono-Ub. From a crystal structure of a complex between UCHL3 and synthetic K27 Ub 2 , we unexpectedly discover that free K27 Ub 2 and K27 Ub 2 -conjugated substrates are natural inhibitors of UCHL3. Using our Ub tools to profile UCHL3's activity, we generate a quantitative kinetic model of the inhibitory mechanism and we find that K27 Ub 2 can inhibit UCHL3 covalently, by binding to its catalytic cysteine, and allosterically, by locking its catalytic loop tightly in place. Based on this inhibition mechanism, we propose that UCHL3 and K27 Ub chains likely sense and regulate each other in cells.
- Department of Cell and Chemical Biology and Oncode Institute, Leiden University Medical Center, Einthovenweg 20, 2333 ZC Leiden, the Netherlands.
Organizational Affiliation: