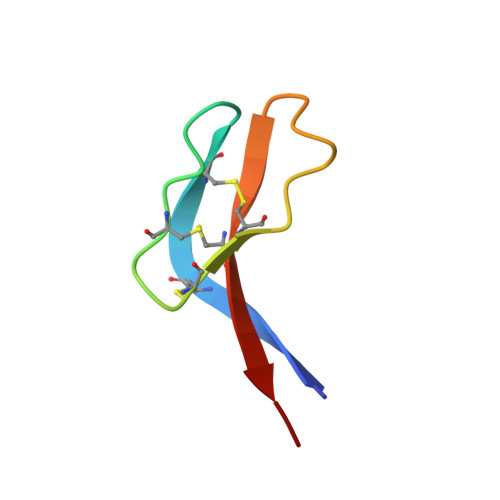
Tick saliva protein Evasin-3 modulates chemotaxis by disrupting CXCL8 interactions with glycosaminoglycans and CXCR2.
Denisov, S.S., Ippel, J.H., Heinzmann, A.C.A., Koenen, R.R., Ortega-Gomez, A., Soehnlein, O., Hackeng, T.M., Dijkgraaf, I.(2019) J Biological Chem 294: 12370-12379
- PubMed: 31235521 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.RA119.008902
- Primary Citation Related Structures:
6QJB - PubMed Abstract:
Chemokines are a group of chemotaxis proteins that regulate cell trafficking and play important roles in immune responses and inflammation. Ticks are blood-sucking parasites that secrete numerous immune-modulatory agents in their saliva to evade host immune responses. Evasin-3 is a small salivary protein that belongs to a class of chemokine-binding proteins isolated from the brown dog tick, Rhipicephalus sanguineus Evasin-3 has been shown to have a high affinity for chemokines CXCL1 and CXCL8 and to diminish inflammation in mice. In the present study, solution NMR spectroscopy was used to investigate the structure of Evasin-3 and its CXCL8-Evasin-3 complex. Evasin-3 is found to disrupt the glycosaminoglycan-binding site of CXCL8 and inhibit the interaction of CXCL8 with CXCR2. Structural data were used to design two novel CXCL8-binding peptides. The linear tEv3 17-56 and cyclic tcEv3 16-56 dPG Evasin-3 variants were chemically synthesized by solid-phase peptide synthesis. The affinity of these newly synthesized variants to CXCL8 was measured by surface plasmon resonance biosensor analysis. The K d values of tEv3 17-56 and tcEv3 16-56 dPG were 27 and 13 nm, respectively. Both compounds effectively inhibited CXCL8-induced migration of polymorphonuclear neutrophils. The present results suggest utility of synthetic Evasin-3 variants as scaffolds for designing and fine-tuning new chemokine-binding agents that suppress immune responses and inflammation.
- Department of Biochemistry, University of Maastricht, Cardiovascular Research Institute Maastricht, 6229 ER, Maastricht, The Netherlands.
Organizational Affiliation: