Polypharmacological Perturbation of the 14-3-3 Adaptor Protein Interactome Stimulates Neurite Outgrowth.
Kaplan, A., Andrei, S.A., van Regteren Altena, A., Simas, T., Banerjee, S.L., Kato, N., Bisson, N., Higuchi, Y., Ottmann, C., Fournier, A.E.(2020) Cell Chem Biol 27: 657-667.e6
- PubMed: 32220335 Search on PubMed
- DOI: https://doi.org/10.1016/j.chembiol.2020.02.010
- Primary Citation Related Structures:
6QDR, 6QDS, 6QDT, 6QDU - PubMed Abstract:
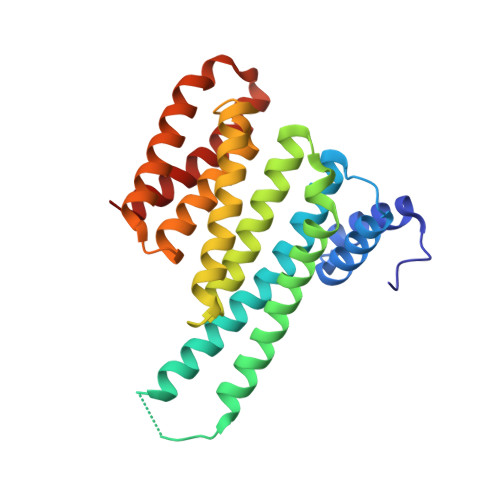
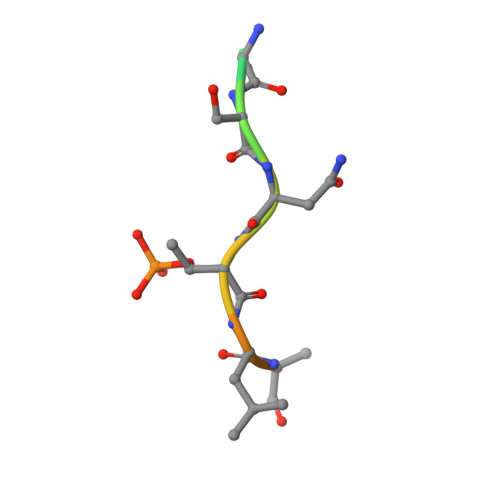
Targeting protein-protein interactions (PPIs) is a promising approach in the development of drugs for many indications. 14-3-3 proteins are a family of phosphoprotein-binding molecules with critical functions in dozens of cell signaling networks. 14-3-3s are abundant in the central nervous system, and the small molecule fusicoccin-A (FC-A), a tool compound that can be used to manipulate 14-3-3 PPIs, enhances neurite outgrowth in cultured neurons. New semisynthetic FC-A derivatives with improved binding affinity for 14-3-3 complexes have recently been developed. Here, we use a series of screens that identify these compounds as potent inducers of neurite outgrowth through a polypharmacological mechanism. Using proteomics and X-ray crystallography, we discover that these compounds extensively regulate the 14-3-3 interactome by stabilizing specific PPIs, while disrupting others. These results provide new insights into the development of drugs to target 14-3-3 PPIs, a potential therapeutic strategy for CNS diseases.
- Department of Neurology and Neurosurgery, Montréal Neurological Institute, McGill University, Montréal, QC, Canada. Electronic address: andrew.kaplan@mail.mcgill.ca.
Organizational Affiliation: