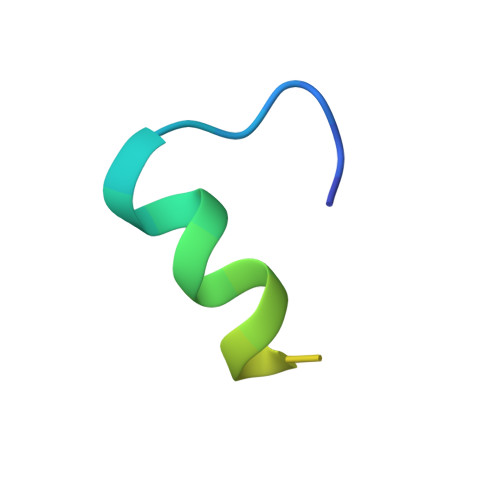
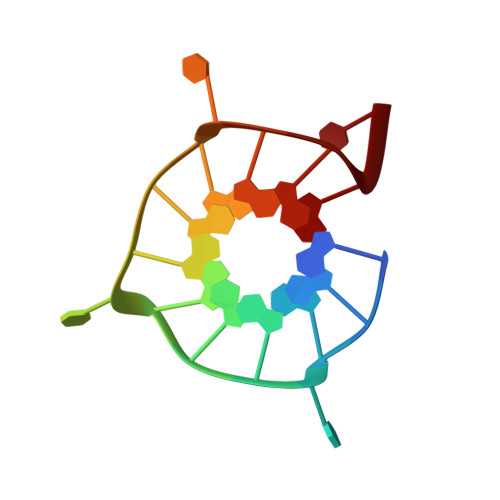
Recognition of different base tetrads by RHAU (DHX36): X-ray crystal structure of the G4 recognition motif bound to the 3'-end tetrad of a DNA G-quadruplex.
Heddi, B., Cheong, V.V., Schmitt, E., Mechulam, Y., Phan, A.T.(2020) J Struct Biol 209: 107399-107399
- PubMed: 31586599 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2019.10.001
- Primary Citation Related Structures:
6Q6R - PubMed Abstract:
G-quadruplexes (G4) are secondary structures of nucleic acids that can form in cells and have diverse biological functions. Several biologically important proteins interact with G-quadruplexes, of which RHAU (or DHX36) - a helicase from the DEAH-box superfamily, was shown to bind and unwind G-quadruplexes efficiently. We report a X-ray co-crystal structure at 1.5 Å resolution of an N-terminal fragment of RHAU bound to an exposed tetrad of a parallel-stranded G-quadruplex. The RHAU peptide folds into an L-shaped α-helix, and binds to a G-quadruplex through π-stacking and electrostatic interactions. X-ray crystal structure of our complex identified key amino acid residues important for G-quadruplex-peptide binding interaction at the 3'-end G•G•G•G tetrad. Together with previous solution and crystal structures of RHAU bound to the 5'-end G•G•G•G and G•G•A•T tetrads, our crystal structure highlights the occurrence of a robust G-quadruplex recognition motif within RHAU that can adapt to different accessible tetrads.
- School of Physical and Mathematical Sciences, Nanyang Technological University, 21 Nanyang Link, Singapore 637371, Singapore; Laboratoire de Biologie et de Pharmacologie Appliquée, CNRS UMR 8113, Ecole Normale Supérieure Paris-Saclay, 61 Avenue du président Wilson, Cachan 94235, France. Electronic address: brahim.heddi@ens-cachan.fr.
Organizational Affiliation: