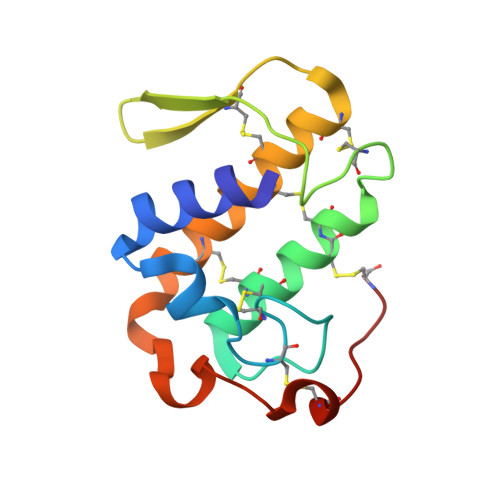
Structural basis for phospholipase A2-like toxin inhibition by the synthetic compound Varespladib (LY315920).
Salvador, G.H.M., Gomes, A.A.S., Bryan-Quiros, W., Fernandez, J., Lewin, M.R., Gutierrez, J.M., Lomonte, B., Fontes, M.R.M.(2019) Sci Rep 9: 17203-17203
- PubMed: 31748642 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-019-53755-5
- Primary Citation Related Structures:
6PWH - PubMed Abstract:
The World Health Organization recently listed snakebite envenoming as a Neglected Tropical Disease, proposing strategies to significantly reduce the global burden of this complex pathology by 2030. In this context, effective adjuvant treatments to complement conventional antivenom therapy based on inhibitory molecules for specific venom toxins have gained renewed interest. Varespladib (LY315920) is a synthetic molecule clinically tested to block inflammatory cascades of several diseases associated with elevated levels of secreted phospholipase A 2 (sPLA 2 ). Most recently, Varespladib was tested against several whole snake venoms and isolated PLA 2 toxins, demonstrating potent inhibitory activity. Herein, we describe the first structural and functional study of the complex between Varespladib and a PLA 2 -like snake venom toxin (MjTX-II). In vitro and in vivo experiments showed this compound's capacity to inhibit the cytotoxic and myotoxic effects of MjTX-II from the medically important South American snake, Bothrops moojeni. Crystallographic and bioinformatics analyses revealed interactions of Varespladib with two specific regions of the toxin, suggesting inhibition occurs by physical blockage of its allosteric activation, preventing the alignment of its functional sites and, consequently, impairing its ability to disrupt membranes. Furthermore, based on the analysis of several crystallographic structures, a distinction between toxin activators and inhibitors is proposed.
- Departamento de Física e Biofísica, Instituto de Biociências, Universidade Estadual Paulista (UNESP), Botucatu, SP, Brazil.
Organizational Affiliation: