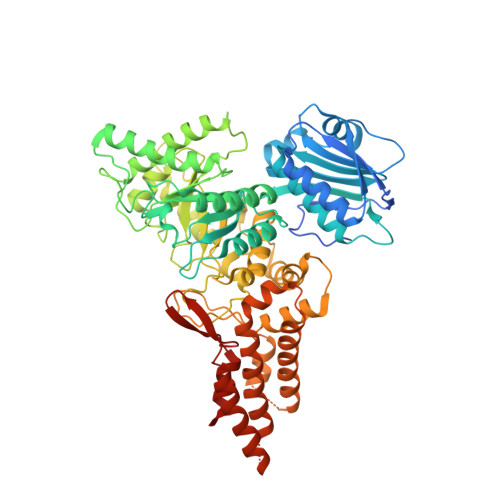
Structural and functional analysis of four family 84 glycoside hydrolases from the opportunistic pathogen Clostridium perfringens.
Pluvinage, B., Massel, P.M., Burak, K., Boraston, A.B.(2019) Glycobiology 30: 49-57
- PubMed: 31701135
- DOI: https://doi.org/10.1093/glycob/cwz069
- Primary Citation of Related Structures:
6PV4, 6PV5, 6PWI - PubMed Abstract:
The opportunistic pathogen Clostridium perfringens possesses the ability to colonize the protective mucin layer in the gastrointestinal tract. To assist this, the C. perfringens genome contains a battery of genes encoding glycoside hydrolases (GHs) that are likely active on mucin glycans, including four genes encoding family 84 GHs: CpGH84A (NagH), CpGH84B (NagI), CpGH84C (NagJ) and CpGH84D (NagK). To probe the potential advantage gained by the expansion of GH84 enzymes in C. perfringens, we undertook the structural and functional characterization of the CpGH84 catalytic modules. Here, we show that these four CpGH84 catalytic modules act as β-N-acetyl-D-glucosaminidases able to hydrolyze N- and O-glycan motifs. CpGH84A and CpGH84D displayed a substrate specificity restricted to terminal β-1,2- and β-1,6-linked N-acetyl-D-glucosamine (GlcNAc). CpGH84B and CpGH84C appear more promiscuous with activity on terminal β-1,2-, β-1,3- and β-1,6-linked GlcNAc; both possess some activity toward β-1,4-linked GlcNAc, but this is dependent upon which monosaccharide it is linked to. Furthermore, all the CpGH84s have different optimum pHs ranging from 5.2 to 7.0. Consistent with their β-N-acetyl-D-glucosaminidase activities, the structures of the four catalytic modules revealed similar folds with a catalytic site including a conserved -1 subsite that binds GlcNAc. However, nonconserved residues in the vicinity of the +1 subsite suggest different accommodation of the sugar preceding the terminal GlcNAc, resulting in subtly different substrate specificities. This structure-function comparison of the four GH84 catalytic modules from C. perfringens reveals their different biochemical properties, which may relate to how they are deployed in the bacterium's niche in the host.
- Biochemistry and Microbiology, University of Victoria, PO Box 3055 STN CSC, Victoria, BC V8W 3P6, Canada.
Organizational Affiliation: