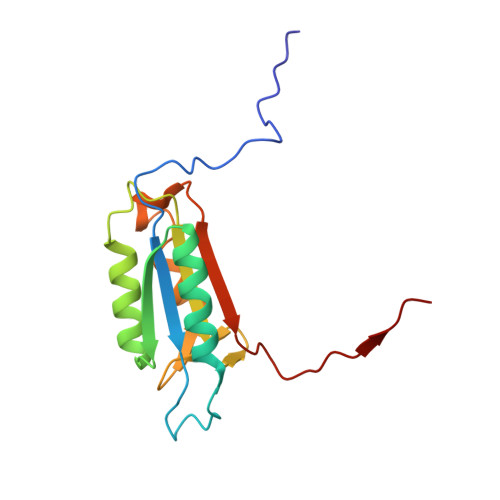
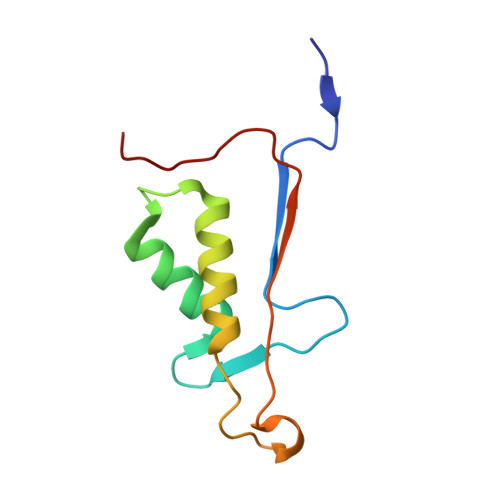
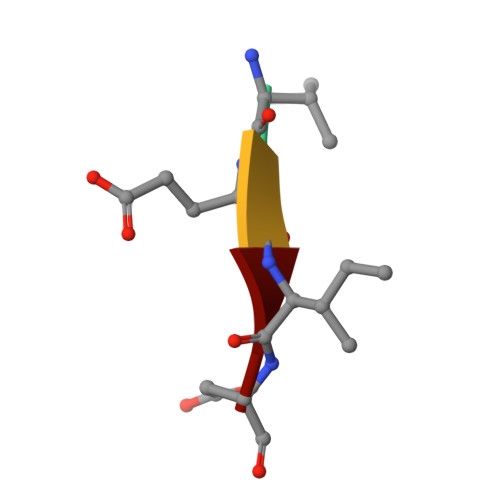
Resurrection of ancestral effector caspases identifies novel networks for evolution of substrate specificity.
Grinshpon, R.D., Shrestha, S., Titus-McQuillan, J., Hamilton, P.T., Swartz, P.D., Clark, A.C.(2019) Biochem J 476: 3475-3492
- PubMed: 31675069 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1042/BCJ20190625
- Primary Citation Related Structures:
6PDQ, 6PPM - PubMed Abstract:
Apoptotic caspases evolved with metazoans more than 950 million years ago (MYA), and a series of gene duplications resulted in two subfamilies consisting of initiator and effector caspases. The effector caspase genes (caspases-3, -6, and -7) were subsequently fixed into the Chordata phylum more than 650 MYA when the gene for a common ancestor (CA) duplicated, and the three effector caspases have persisted throughout mammalian evolution. All caspases prefer an aspartate residue at the P1 position of substrates, so each caspase evolved discrete cellular roles through changes in substrate recognition at the P4 position combined with allosteric regulation. We examined the evolution of substrate specificity in caspase-6, which prefers valine at the P4 residue, compared with caspases-3 and -7, which prefer aspartate, by reconstructing the CA of effector caspases (AncCP-Ef1) and the CA of caspase-6 (AncCP-6An). We show that AncCP-Ef1 is a promiscuous enzyme with little distinction between Asp, Val, or Leu at P4. The specificity of caspase-6 was defined early in its evolution, where AncCP-6An demonstrates a preference for Val over Asp at P4. Structures of AncCP-Ef1 and of AncCP-6An show a network of charged amino acids near the S4 pocket that, when combined with repositioning a flexible active site loop, resulted in a more hydrophobic binding pocket in AncCP-6An. The ancestral protein reconstructions show that the caspase-hemoglobinase fold has been conserved for over 650 million years and that only three substitutions in the scaffold are necessary to shift substrate selection toward Val over Asp.
- Department of Biology, University of Texas at Arlington, Arlington, TX 76019, U.S.A.
Organizational Affiliation: