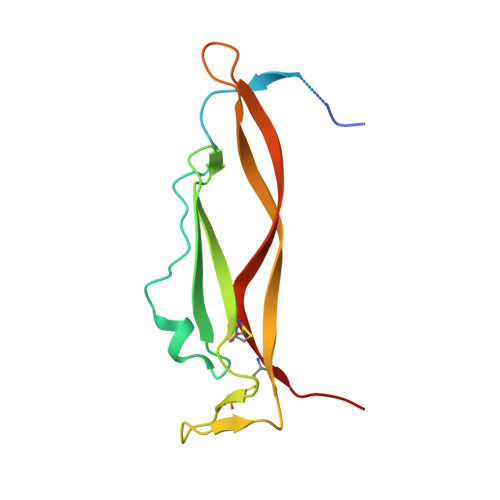
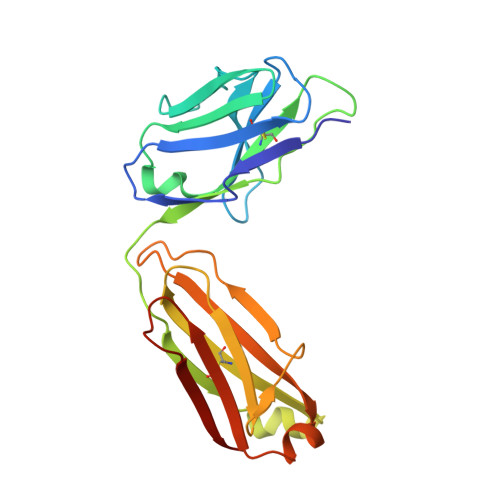
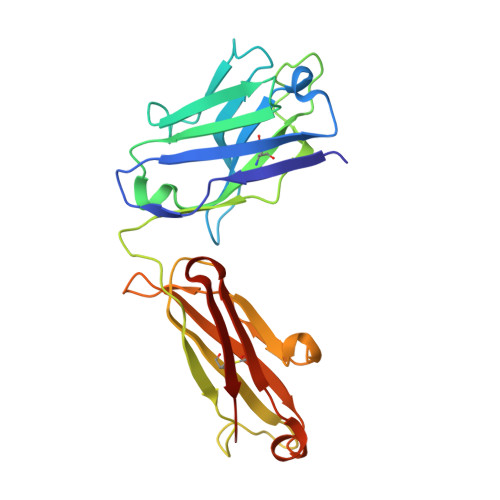
Dissecting the molecular basis of high viscosity of monospecific and bispecific IgG antibodies.
Tilegenova, C., Izadi, S., Yin, J., Huang, C.S., Wu, J., Ellerman, D., Hymowitz, S.G., Walters, B., Salisbury, C., Carter, P.J.(null) MAbs 12: 1692764-1692764
- PubMed: 31779513 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1080/19420862.2019.1692764
- Primary Citation Related Structures:
6PPG - PubMed Abstract:
Some antibodies exhibit elevated viscosity at high concentrations, making them poorly suited for therapeutic applications requiring administration by injection such as subcutaneous or ocular delivery. Here we studied an anti-IL-13/IL-17 bispecific IgG 4 antibody, which has anomalously high viscosity compared to its parent monospecific antibodies. The viscosity of the bispecific IgG 4 in solution was decreased by only ~30% in the presence of NaCl, suggesting electrostatic interactions are insufficient to fully explain the drivers of viscosity. Intriguingly, addition of arginine-HCl reduced the viscosity of the bispecific IgG 4 by ~50% to its parent IgG level. These data suggest that beyond electrostatics, additional types of interactions such as cation-π and/or π-π may contribute to high viscosity more significantly than previously understood. Molecular dynamics simulations of antibody fragments in the mixed solution of free arginine and explicit water were conducted to identify hotspots involved in self-interactions. Exposed surface aromatic amino acids displayed an increased number of contacts with arginine. Mutagenesis of the majority of aromatic residues pinpointed by molecular dynamics simulations effectively decreased the solution's viscosity when tested experimentally. This mutational method to reduce the viscosity of a bispecific antibody was extended to a monospecific anti-GCGR IgG 1 antibody with elevated viscosity. In all cases, point mutants were readily identified that both reduced viscosity and retained antigen-binding affinity. These studies demonstrate a new approach to mitigate high viscosity of some antibodies by mutagenesis of surface-exposed aromatic residues on complementarity-determining regions that may facilitate some clinical applications.
- Antibody Engineering, Genentech Inc., South San Francisco, CA, USA.
Organizational Affiliation: