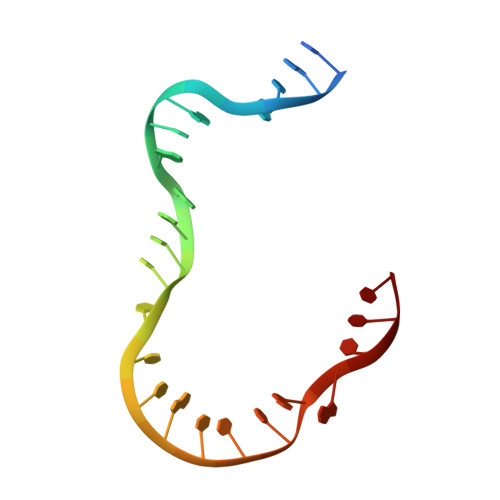
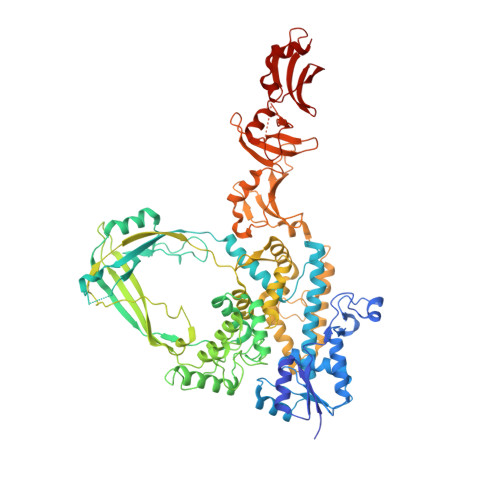
Mechanistic insights from structure of Mycobacterium smegmatis topoisomerase I with ssDNA bound to both N- and C-terminal domains.
Cao, N., Tan, K., Zuo, X., Annamalai, T., Tse-Dinh, Y.C.(2020) Nucleic Acids Res 48: 4448-4462
- PubMed: 32232337 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkaa201
- Primary Citation Related Structures:
6PCM - PubMed Abstract:
Type IA topoisomerases interact with G-strand and T-strand ssDNA to regulate DNA topology. However, simultaneous binding of two ssDNA segments to a type IA topoisomerase has not been observed previously. We report here the crystal structure of a type IA topoisomerase with ssDNA segments bound in opposite polarity to the N- and C-terminal domains. Titration of small ssDNA oligonucleotides to Mycobacterium smegmatis topoisomerase I with progressive C-terminal deletions showed that the C-terminal region has higher affinity for ssDNA than the N-terminal active site. This allows the C-terminal domains to capture one strand of underwound negatively supercoiled DNA substrate first and position the N-terminal domains to bind and cleave the opposite strand in the relaxation reaction. Efficiency of negative supercoiling relaxation increases with the number of domains that bind ssDNA primarily with conserved aromatic residues and possibly with assistance from polar/basic residues. A comparison of bacterial topoisomerase I structures showed that a conserved transesterification unit (N-terminal toroid structure) for cutting and rejoining of a ssDNA strand can be combined with two different types of C-terminal ssDNA binding domains to form diverse bacterial topoisomerase I enzymes that are highly efficient in their physiological role of preventing excess negative supercoiling in the genome.
- Department of Chemistry and Biochemistry, Florida International University, Miami, FL 33199, USA.
Organizational Affiliation: