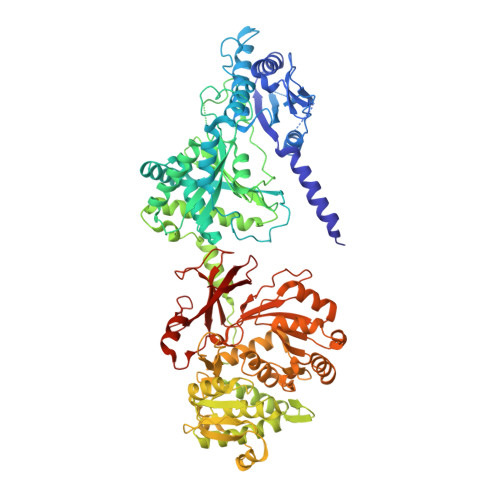
Structures of teixobactin-producing nonribosomal peptide synthetase condensation and adenylation domains.
Tan, K., Zhou, M., Jedrzejczak, R.P., Wu, R., Higuera, R.A., Borek, D., Babnigg, G., Joachimiak, A.(2020) Curr Res Struct Biol 2: 14-24
- PubMed: 34235466 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.crstbi.2020.01.002
- Primary Citation Related Structures:
6OYF, 6OZV, 6P1J, 6P3I, 6P4U - PubMed Abstract:
The recently discovered antibiotic teixobactin is produced by uncultured soil bacteria. The antibiotic inhibits cell wall synthesis of Gram-positive bacteria by binding to precursors of cell wall building blocks, and therefore it is thought to be less vulnerable to development of resistance. Teixobactin is synthesized by two nonribosomal peptide synthetases (NRPSs), encoded by txo1 and txo2 genes. Like other NRPSs, the Txo1 and Txo2 synthetases are large, multifunctional, and comprised of several modules. Each module is responsible for catalysis of a distinct step of teixobactin synthesis and contains specific functional units, commonly including a condensation (C) domain, an adenylation (A) domain, and a peptidyl carrier protein (PCP) domain. Here we report the structures of the C-A bidomains of the two L-Ser condensing modules, from Txo1 and Txo2, respectively. In the structure of the C domain of the L-Ser subunit of Txo1, a large conformational change is observed, featuring an outward swing of its N-terminal α-helix. This repositioning, if functionally validated, provides the necessary conformational change for the condensation reaction in C domain, and likely represents a regulatory mechanism. In an A core subdomain, a well-coordinated Mg 2+ cation is observed, which is required in the adenylation reaction. The Mg 2+ -binding site is defined by a largely conserved amino acid sequence motif and is coordinated by the α-phosphate group of AMP (or ATP) when present, providing some structural evidence for the role of the metal cation in the catalysis of A domain.
- Center for Structural Genomics of Infectious Diseases, University of Chicago, 5735 South Ellis Avenue, Chicago, IL 60637, USA.
Organizational Affiliation: