A Strain-Specific Inhibitor of Receptor-Bound HIV-1 Targets a Pocket near the Fusion Peptide.
Ozorowski, G., Torres, J.L., Santos-Martins, D., Forli, S., Ward, A.B.(2020) Cell Rep 33: 108428-108428
- PubMed: 33238117 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.celrep.2020.108428
- Primary Citation Related Structures:
6OPN, 6OPO, 6OPP, 6OPQ, 6X5B, 6X5C - PubMed Abstract:
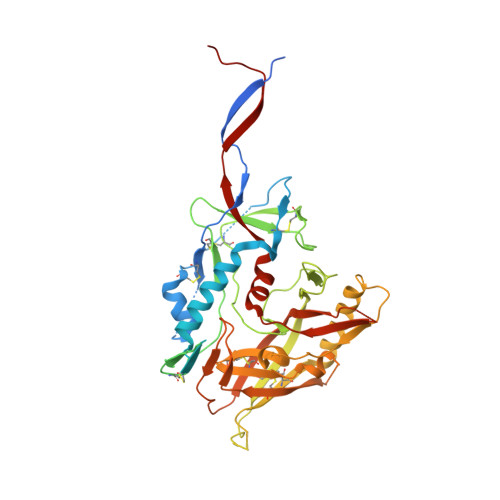
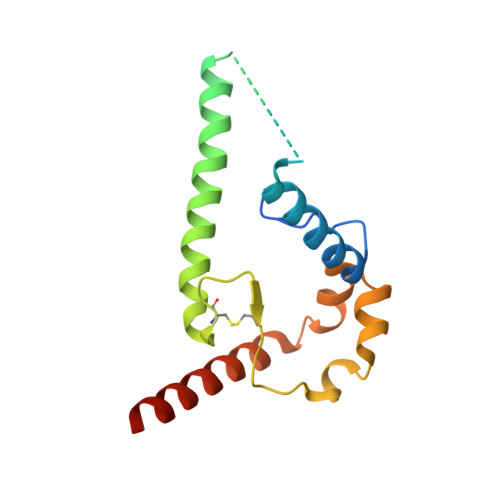
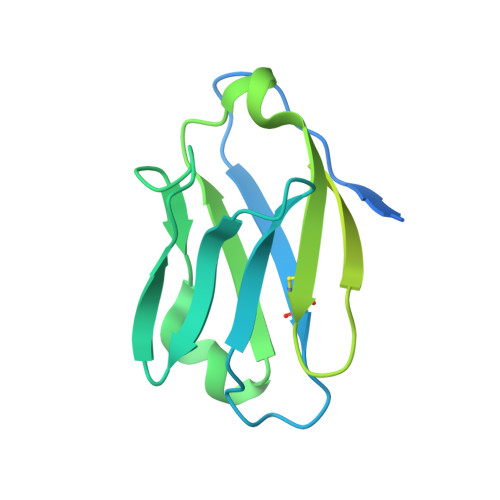
Disruption of viral fusion represents a viable, albeit under-explored, target for HIV therapeutics. Here, while studying the receptor-bound envelope glycoprotein conformation by cryoelectron microscopy (cryo-EM), we identify a pocket near the base of the trimer containing a bound detergent molecule and perform in silico drug screening by using a library of drug-like and commercially available molecules. After down-selection, we solve cryo-EM structures that validate the binding of two small molecule hits in very similar manners to the predicted binding poses, including interactions with aromatic residues within the fusion peptide. One of the molecules demonstrates low micromolar inhibition of the autologous virus by using a very rare phenylalanine in the fusion peptide and stabilizing the surrounding region. This work demonstrates that small molecules can target the fusion process, providing an additional target for anti-HIV therapeutics, and highlights the need to explore how fusion peptide sequence variations affect receptor-mediated conformational states across diverse HIV strains.
- Department of Integrative Structural and Computational Biology, The Scripps Research Institute, La Jolla, CA 92037, USA; Center for HIV/AIDS Vaccine Immunology and Immunogen Discovery, International AIDS Vaccine Initiative Neutralizing Antibody Center, and Collaboration for AIDS Vaccine Discovery, The Scripps Research Institute, La Jolla, CA 92037, USA.
Organizational Affiliation: