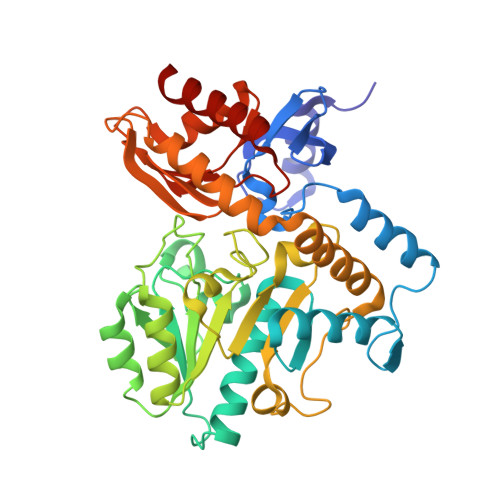
Mechanism of Inactivation of Ornithine Aminotransferase by (1S,3S)-3-Amino-4-(hexafluoropropan-2-ylidenyl)cyclopentane-1-carboxylic Acid.
Moschitto, M.J., Doubleday, P.F., Catlin, D.S., Kelleher, N.L., Liu, D., Silverman, R.B.(2019) J Am Chem Soc 141: 10711-10721
- PubMed: 31251613 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/jacs.9b03254
- Primary Citation Related Structures:
6OIA - PubMed Abstract:
The inhibition of ornithine aminotransferase (OAT), a pyridoxal 5'-phosphate-dependent enzyme, has been implicated as a treatment for hepatocellular carcinoma (HCC), the most common form of liver cancer, for which there is no effective treatment. From a previous evaluation of our aminotransferase inhibitors, (1 S ,3 S )-3-amino-4-(perfluoropropan-2-ylidene)cyclopentane-1-carboxylic acid hydrochloride ( 1 ) was found to be a selective and potent inactivator of human OAT ( h OAT), which inhibited the growth of HCC in athymic mice implanted with human-derived HCC, even at a dose of 0.1 mg/kg. Currently, investigational new drug (IND)-enabling studies with 1 are underway. The inactivation mechanism of 1 , however, has proved to be elusive. Here we propose three possible mechanisms, based on mechanisms of known aminotransferase inactivators: Michael addition, enamine addition, and fluoride ion elimination followed by conjugate addition. On the basis of crystallography and intact protein mass spectrometry, it was determined that 1 inactivates h OAT through fluoride ion elimination to an activated 1,1'-difluoroolefin, followed by conjugate addition and hydrolysis. This result was confirmed with additional studies, including the detection of the cofactor structure by mass spectrometry and through the identification of turnover metabolites. On the basis of this inactivation mechanism and to provide further evidence for the mechanism, analogues of 1 ( 19 , 20 ) were designed, synthesized, and demonstrated to have the predicted selective inactivation mechanism. These analogues highlight the importance of the trifluoromethyl group and provide a basis for future inactivator design.
- Department of Chemistry and Biochemistry , Loyola University Chicago , Chicago , Illinois 60660 , United States.
Organizational Affiliation: