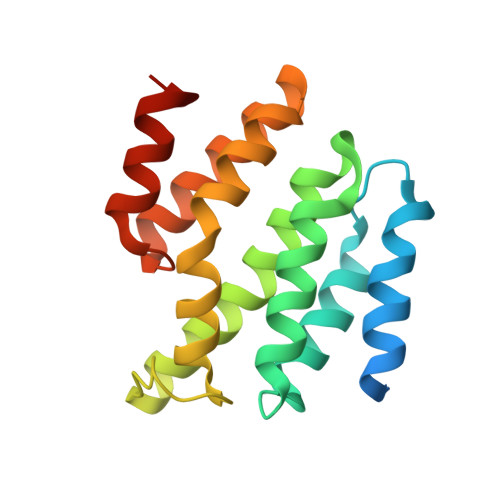
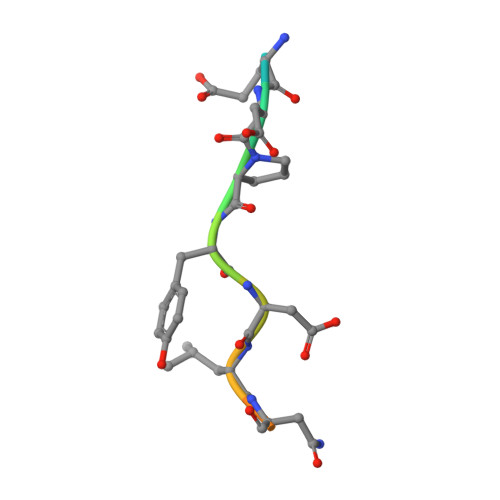
Identification of Three Sequence Motifs in the Transcription Termination Factor Sen1 that Mediate Direct Interactions with Nrd1.
Zhang, Y., Chun, Y., Buratowski, S., Tong, L.(2019) Structure 27: 1156-1161.e4
- PubMed: 31104813
- DOI: https://doi.org/10.1016/j.str.2019.04.005
- Primary Citation of Related Structures:
6O3W, 6O3X, 6O3Y - PubMed Abstract:
The Nrd1-Nab3-Sen1 (NNS) complex carries out the transcription termination of non-coding RNAs (ncRNAs) by RNA polymerase II (Pol II) in yeast, although the detailed interactions among its subunits remain obscure. Here we have identified three sequence motifs in Sen1 that mediate direct interactions with the Pol II CTD interaction domain (CID) of Nrd1, determined the crystal structures of these Nrd1 interaction motifs (NIMs) bound to the CID, and characterized the interactions in vitro and in yeast. Removal of all three NIMs abolishes NNS complex formation and gives rise to ncRNA termination defects.
- Department of Biological Sciences, Columbia University, New York, NY 10027, USA.
Organizational Affiliation: