KLK4 Inhibition by Cyclic and Acyclic Peptides: Structural and Dynamical Insights into Standard-Mechanism Protease Inhibitors.
Riley, B.T., Ilyichova, O., de Veer, S.J., Swedberg, J.E., Wilson, E., Hoke, D.E., Harris, J.M., Buckle, A.M.(2019) Biochemistry 58: 2524-2533
- PubMed: 31058493 Search on PubMed
- DOI: https://doi.org/10.1021/acs.biochem.9b00191
- Primary Citation Related Structures:
4KEL, 6O21 - PubMed Abstract:
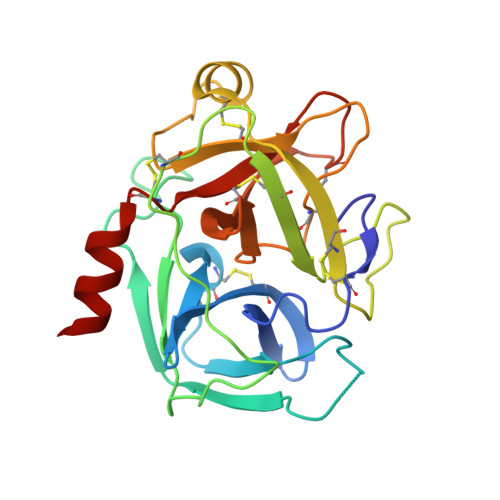
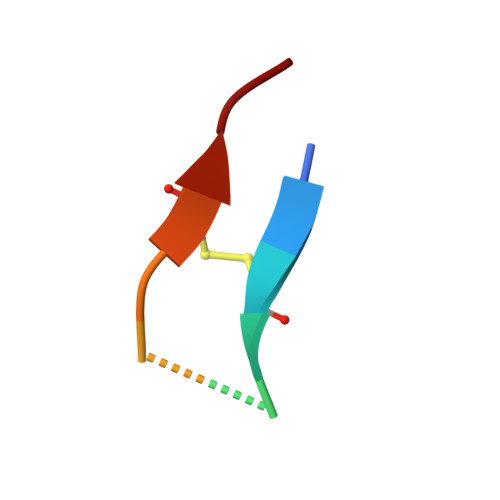
Sunflower trypsin inhibitor (SFTI-1) is a 14 amino acid serine protease inhibitor. The dual antiparallel β-sheet arrangement of SFTI-1 is stabilized by an N-terminal-C-terminal backbone cyclization and a further disulfide bridge to form a final bicyclic structure. This constrained structure is further rigidified by an extensive network of internal hydrogen bonds. Thus, the structure of SFTI-1 in solution resembles the protease-bound structure, reducing the entropic penalty upon protease binding. When cleaved at the scissile bond, it is thought that the rigidifying features of SFTI-1 maintain its structure, allowing the scissile bond to be reformed. The lack of structural plasticity for SFTI-1 is proposed to favor initial protease binding and continued occupancy in the protease active site, resulting in an equilibrium between the cleaved and uncleaved inhibitor in the presence of a protease. We have determined, at 1.15 Å resolution, the X-ray crystal structures of complexes between human kallikrein-related peptidase 4 (KLK4) and SFTI-FCQR(Asn14) and between KLK4 and an acyclic form of the same inhibitor, SFTI-FCQR(Asn14)[1,14], with the latter displaying a cleaved scissile bond. Structural analysis and MD simulations together reveal the roles of the altered contact sequence, intramolecular hydrogen bonding network, and backbone cyclization in altering the state of SFTI's scissile bond ligation at the protease active site. Taken together, the data presented reveal insights into the role of dynamics in the standard-mechanism inhibition and suggest that modifications on the non-contact strand may be a useful, underexplored approach for generating further potent or selective SFTI-based inhibitors against members of the serine protease family.
- Department of Biochemistry and Molecular Biology, Biomedicine Discovery Institute , Monash University , Clayton , Victoria 3800 , Australia.
Organizational Affiliation: