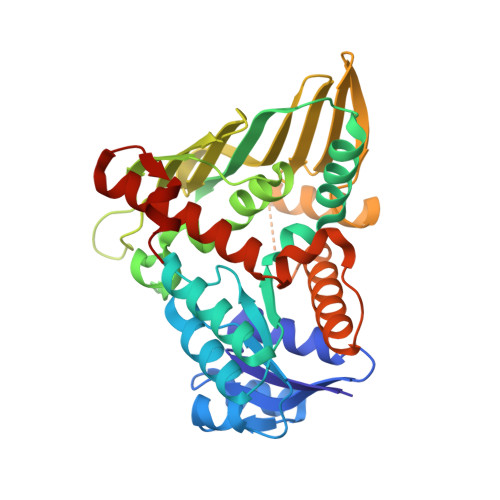
On the structure and function of Escherichia coli YjhC: An oxidoreductase involved in bacterial sialic acid metabolism.
Horne, C.R., Kind, L., Davies, J.S., Dobson, R.C.J.(2020) Proteins 88: 654-668
- PubMed: 31697432
- DOI: https://doi.org/10.1002/prot.25846
- Primary Citation Related Structures:
6O15 - PubMed Abstract:
Human pathogenic and commensal bacteria have evolved the ability to scavenge host-derived sialic acids and subsequently degrade them as a source of nutrition. Expression of the Escherichia coli yjhBC operon is controlled by the repressor protein nanR, which regulates the core machinery responsible for the import and catabolic processing of sialic acid. The role of the yjhBC encoded proteins is not known-here, we demonstrate that the enzyme YjhC is an oxidoreductase/dehydrogenase involved in bacterial sialic acid degradation. First, we demonstrate in vivo using knockout experiments that YjhC is broadly involved in carbohydrate metabolism, including that of N-acetyl-d-glucosamine, N-acetyl-d-galactosamine and N-acetylneuraminic acid. Differential scanning fluorimetry demonstrates that YjhC binds N-acetylneuraminic acid and its lactone variant, along with NAD(H), which is consistent with its role as an oxidoreductase. Next, we solved the crystal structure of YjhC in complex with the NAD(H) cofactor to 1.35 Å resolution. The protein fold belongs to the Gfo/Idh/MocA protein family. The dimeric assembly observed in the crystal form is confirmed through solution studies. Ensemble refinement reveals a flexible loop region that may play a key role during catalysis, providing essential contacts to stabilize the substrate-a unique feature to YjhC among closely related structures. Guided by the structure, in silico docking experiments support the binding of sialic acid and several common derivatives in the binding pocket, which has an overall positive charge distribution. Taken together, our results verify the role of YjhC as a bona fide oxidoreductase/dehydrogenase and provide the first evidence to support its involvement in sialic acid metabolism.
- Biomolecular Interaction Centre and School of Biological Sciences, University of Canterbury, Christchurch, New Zealand.
Organizational Affiliation: