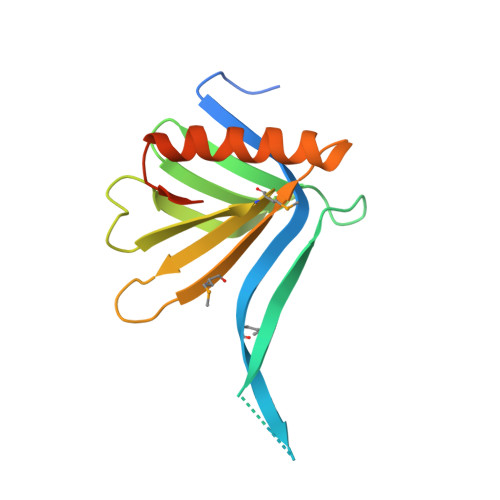
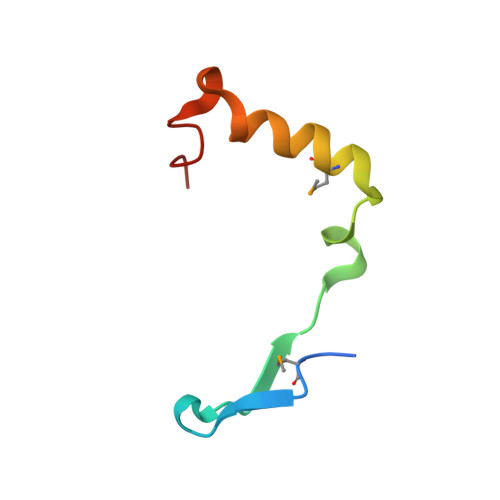
REC114 Partner ANKRD31 Controls Number, Timing, and Location of Meiotic DNA Breaks.
Boekhout, M., Karasu, M.E., Wang, J., Acquaviva, L., Pratto, F., Brick, K., Eng, D.Y., Xu, J., Camerini-Otero, R.D., Patel, D.J., Keeney, S.(2019) Mol Cell 74: 1053-1068.e8
- PubMed: 31003867 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molcel.2019.03.023
- Primary Citation Related Structures:
6NXF - PubMed Abstract:
Double-strand breaks (DSBs) initiate the homologous recombination that is crucial for meiotic chromosome pairing and segregation. Here, we unveil mouse ANKRD31 as a lynchpin governing multiple aspects of DSB formation. Spermatocytes lacking ANKRD31 have altered DSB locations and fail to target DSBs to the pseudoautosomal regions (PARs) of sex chromosomes. They also have delayed and/or fewer recombination sites but, paradoxically, more DSBs, suggesting DSB dysregulation. Unrepaired DSBs and pairing failures-stochastic on autosomes, nearly absolute on X and Y-cause meiotic arrest and sterility in males. Ankrd31-deficient females have reduced oocyte reserves. A crystal structure defines a pleckstrin homology (PH) domain in REC114 and its direct intermolecular contacts with ANKRD31. In vivo, ANKRD31 stabilizes REC114 association with the PAR and elsewhere. Our findings inform a model in which ANKRD31 is a scaffold anchoring REC114 and other factors to specific genomic locations, thereby regulating DSB formation.
- Molecular Biology Program, Memorial Sloan Kettering Cancer Center, New York, NY 10065, USA.
Organizational Affiliation: