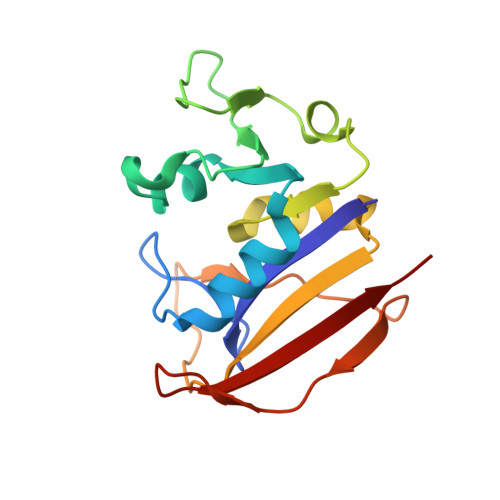
Crystal structures of the closed form of Mycobacterium tuberculosis dihydrofolate reductase in complex with dihydrofolate and antifolates.
Ribeiro, J.A., Chavez-Pacheco, S.M., de Oliveira, G.S., Silva, C.D.S., Giudice, J.H.P., Libreros-Zuniga, G.A., Dias, M.V.B.(2019) Acta Crystallogr D Struct Biol 75: 682-693
- PubMed: 31282477 Search on PubMed
- DOI: https://doi.org/10.1107/S205979831900901X
- Primary Citation Related Structures:
6NNC, 6NND, 6NNE, 6NNH, 6NNI - PubMed Abstract:
Tuberculosis is a disease caused by Mycobacterium tuberculosis and is the leading cause of death from a single infectious pathogen, with a high prevalence in developing countries in Africa and Asia. There still is a need for the development or repurposing of novel therapies to combat this disease owing to the long-term nature of current therapies and because of the number of reported resistant strains. Here, structures of dihydrofolate reductase from M. tuberculosis (MtDHFR), which is a key target of the folate pathway, are reported in complex with four antifolates, pyrimethamine, cycloguanil, diaverdine and pemetrexed, and its substrate dihydrofolate in order to understand their binding modes. The structures of all of these complexes were obtained in the closed-conformation state of the enzyme and a fine structural analysis indicated motion in key regions of the substrate-binding site and different binding modes of the ligands. In addition, the affinities, through K d measurement, of diaverdine and methotrexate have been determined; MtDHFR has a lower affinity (highest K d ) for diaverdine than pyrimethamine and trimethoprim, and a very high affinity for methotrexate, as expected. The structural comparisons and analysis described in this work provide new information about the plasticity of MtDHFR and the binding effects of different antifolates.
- Department of Microbiology, Institute of Biomedical Science, University of São Paulo, Avenida Professor Lineu Prestes 1374, São Paulo 05508-000, Brazil.
Organizational Affiliation: