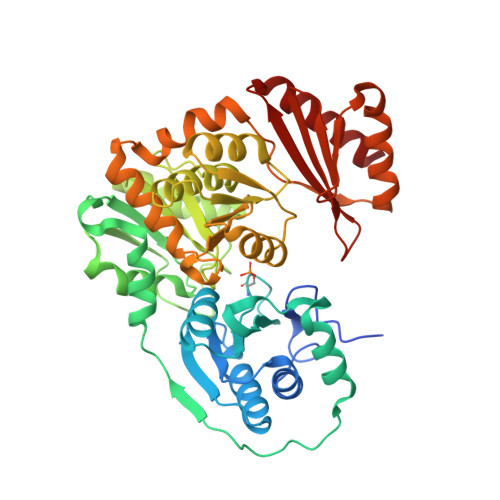
Structural and dynamical description of the enzymatic reaction of a phosphohexomutase.
Stiers, K.M., Graham, A.C., Zhu, J.S., Jakeman, D.L., Nix, J.C., Beamer, L.J.(2019) Struct Dyn 6: 024703-024703
- PubMed: 31041362
- DOI: https://doi.org/10.1063/1.5092803
- Primary Citation Related Structures:
6NN1, 6NN2, 6NNN, 6NNO, 6NNP, 6NNS, 6NNT, 6NNU, 6NOL, 6NOQ, 6NP8, 6NPX, 6NQE, 6NQF, 6NQG - PubMed Abstract:
Enzymes are known to adopt various conformations at different points along their catalytic cycles. Here, we present a comprehensive analysis of 15 isomorphous, high resolution crystal structures of the enzyme phosphoglucomutase from the bacterium Xanthomonas citri . The protein was captured in distinct states critical to function, including enzyme-substrate, enzyme-product, and enzyme-intermediate complexes. Key residues in ligand recognition and regions undergoing conformational change are identified and correlated with the various steps of the catalytic reaction. In addition, we use principal component analysis to examine various subsets of these structures with two goals: (1) identifying sites of conformational heterogeneity through a comparison of room temperature and cryogenic structures of the apo-enzyme and (2) a priori clustering of the enzyme-ligand complexes into functionally related groups, showing sensitivity of this method to structural features difficult to detect by traditional methods. This study captures, in a single system, the structural basis of diverse substrate recognition, the subtle impact of covalent modification, and the role of ligand-induced conformational change in this representative enzyme of the α-D-phosphohexomutase superfamily.
- Biochemistry Department, University of Missouri, 117 Schweitzer Hall, Columbia, Missouri 65211, USA.
Organizational Affiliation: