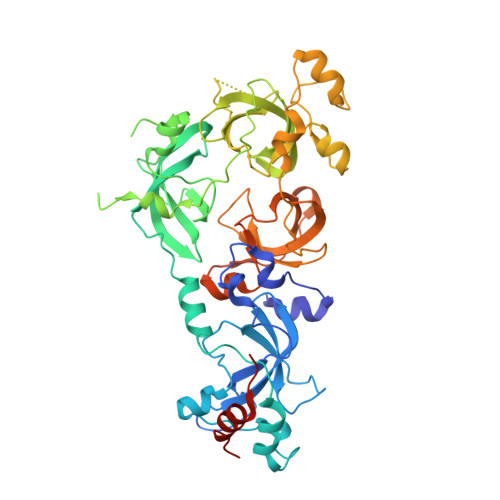
Structural Basis for EPC1-Mediated Recruitment of MBTD1 into the NuA4/TIP60 Acetyltransferase Complex.
Zhang, H., Devoucoux, M., Song, X., Li, L., Ayaz, G., Cheng, H., Tempel, W., Dong, C., Loppnau, P., Cote, J., Min, J.(2020) Cell Rep 30: 3996-4002.e4
- PubMed: 32209463 Search on PubMed
- DOI: https://doi.org/10.1016/j.celrep.2020.03.003
- Primary Citation Related Structures:
6NFX - PubMed Abstract:
MBTD1, a H4K20me reader, has recently been identified as a component of the NuA4/TIP60 acetyltransferase complex, regulating gene expression and DNA repair. NuA4/TIP60 inhibits 53BP1 binding to chromatin through recognition of the H4K20me mark by MBTD1 and acetylation of H2AK15, blocking the ubiquitination mark required for 53BP1 localization at DNA breaks. The NuA4/TIP60 non-catalytic subunit EPC1 enlists MBTD1 into the complex, but the detailed molecular mechanism remains incompletely explored. Here, we present the crystal structure of the MBTD1-EPC1 complex, revealing a hydrophobic C-terminal fragment of EPC1 engaging the MBT repeats of MBTD1 in a site distinct from the H4K20me binding site. Different cellular assays validate the physiological significance of the key residues involved in the MBTD1-EPC1 interaction. Our study provides a structural framework for understanding the mechanism by which MBTD1 recruits the NuA4/TIP60 acetyltransferase complex to influence transcription and DNA repair pathway choice.
- Structural Genomics Consortium, University of Toronto, Toronto, ON M5G 1L7, Canada.
Organizational Affiliation: