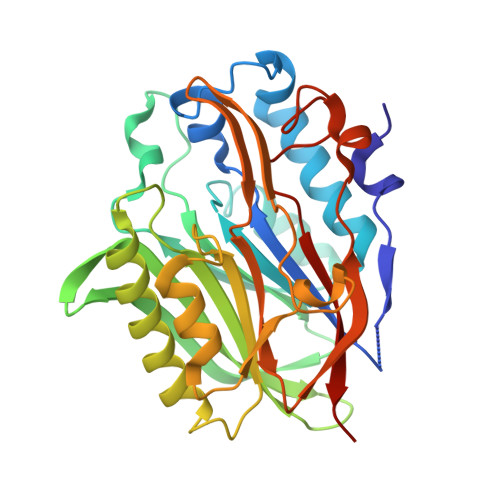
The metabolites NADP+and NADPH are the targets of the circadian protein Nocturnin (Curled).
Estrella, M.A., Du, J., Chen, L., Rath, S., Prangley, E., Chitrakar, A., Aoki, T., Schedl, P., Rabinowitz, J., Korennykh, A.(2019) Nat Commun 10: 2367-2367
- PubMed: 31147539 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-019-10125-z
- Primary Citation Related Structures:
6NF0 - PubMed Abstract:
Nocturnin (NOCT) is a rhythmically expressed protein that regulates metabolism under the control of circadian clock. It has been proposed that NOCT deadenylates and regulates metabolic enzyme mRNAs. However, in contrast to other deadenylases, purified NOCT lacks the deadenylase activity. To identify the substrate of NOCT, we conducted a mass spectrometry screen and report that NOCT specifically and directly converts the dinucleotide NADP + into NAD + and NADPH into NADH. Further, we demonstrate that the Drosophila NOCT ortholog, Curled, has the same enzymatic activity. We obtained the 2.7 Å crystal structure of the human NOCT•NADPH complex, which revealed that NOCT recognizes the chemically unique ribose-phosphate backbone of the metabolite, placing the 2'-terminal phosphate productively for removal. We provide evidence for NOCT targeting to mitochondria and propose that NADP(H) regulation, which takes place at least in part in mitochondria, establishes the molecular link between circadian clock and metabolism.
- 216 Schultz Laboratory, Department of Molecular Biology, Princeton, NJ, 08544, USA.
Organizational Affiliation: