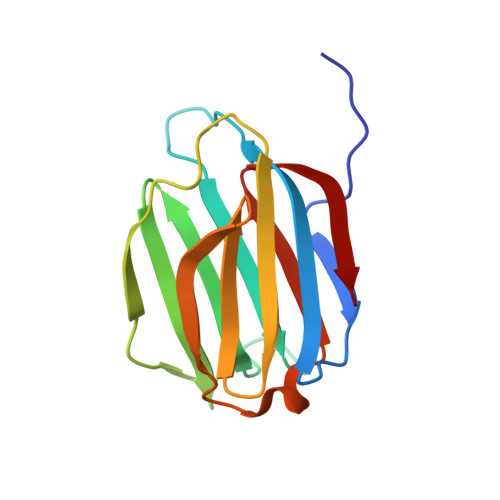
The oligomeric assembly of galectin-11 is critical for anti-parasitic activity in sheep (Ovis aries).
Sakthivel, D., Preston, S., Gasser, R.B., Costa, T.P.S.D., Hernandez, J.N., Shahine, A., Shakif-Azam, M.D., Lock, P., Rossjohn, J., Perugini, M.A., Gonzalez, J.F., Meeusen, E., Piedrafita, D., Beddoe, T.(2020) Commun Biol 3: 464-464
- PubMed: 32826940 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s42003-020-01179-7
- Primary Citation Related Structures:
6N44 - PubMed Abstract:
Galectins are a family of glycan-binding molecules with a characteristic affinity for ß-D-glycosides that mediate a variety of important cellular functions, including immune and inflammatory responses. Galectin-11 (LGALS-11) has been recently identified as a mediator induced specifically in animals against gastrointestinal nematodes and can interfere with parasite growth and development. Here, we report that at least two natural genetic variants of LGALS-11 exist in sheep, and demonstrate fundamental differences in anti-parasitic activity, correlated with their ability to dimerise. This study improves our understanding of the role of galectins in the host immune and inflammatory responses against parasitic nematodes and provides a basis for genetic studies toward selective breeding of animals for resistance to parasites.
- Department of Biochemistry and Molecular Biology, Monash University, Clayton, VIC, 3800, Australia.
Organizational Affiliation: