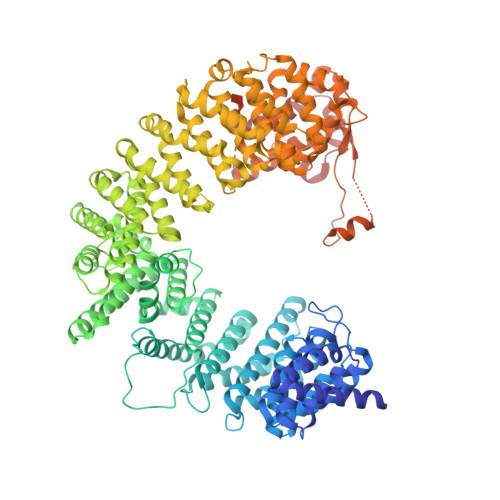
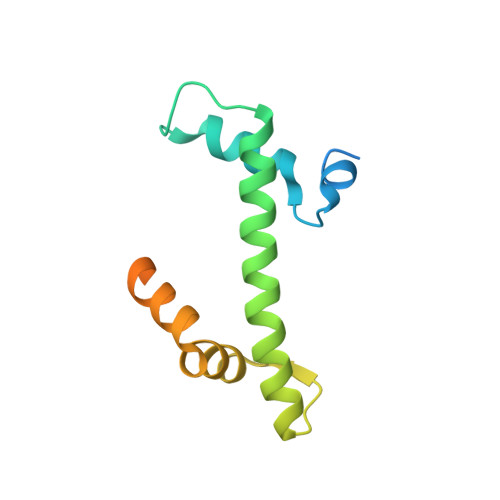
Importin-9 wraps around the H2A-H2B core to act as nuclear importer and histone chaperone.
Padavannil, A., Sarkar, P., Kim, S.J., Cagatay, T., Jiou, J., Brautigam, C.A., Tomchick, D.R., Sali, A., D'Arcy, S., Chook, Y.M.(2019) Elife 8
- PubMed: 30855230 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.7554/eLife.43630
- Primary Citation Related Structures:
6N1Z - PubMed Abstract:
We report the crystal structure of nuclear import receptor Importin-9 bound to its cargo, the histones H2A-H2B. Importin-9 wraps around the core, globular region of H2A-H2B to form an extensive interface. The nature of this interface coupled with quantitative analysis of deletion mutants of H2A-H2B suggests that the NLS-like sequences in the H2A-H2B tails play a minor role in import. Importin-9•H2A-H2B is reminiscent of interactions between histones and histone chaperones in that it precludes H2A-H2B interactions with DNA and H3-H4 as seen in the nucleosome. Like many histone chaperones, which prevent inappropriate non-nucleosomal interactions, Importin-9 also sequesters H2A-H2B from DNA. Importin-9 appears to act as a storage chaperone for H2A-H2B while escorting it to the nucleus. Surprisingly, RanGTP does not dissociate Importin-9•H2A-H2B but assembles into a RanGTP•Importin-9•H2A-H2B complex. The presence of Ran in the complex, however, modulates Imp9-H2A-H2B interactions to facilitate its dissociation by DNA and assembly into a nucleosome.
- Department of Pharmacology, University of Texas Southwestern Medical Center, Dallas, United States.
Organizational Affiliation: