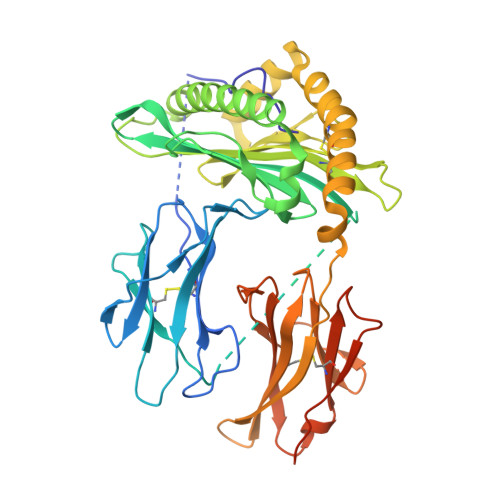
Altered Binding of Tumor Antigenic Peptides to MHC Class I Affects CD8+T Cell-Effector Responses.
Clancy-Thompson, E., Devlin, C.A., Tyler, P.M., Servos, M.M., Ali, L.R., Ventre, K.S., Bhuiyan, M.A., Bruck, P.T., Birnbaum, M.E., Dougan, S.K.(2018) Cancer Immunol Res 6: 1524-1536
- PubMed: 30352798 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1158/2326-6066.CIR-18-0348
- Primary Citation Related Structures:
6MP0, 6MP1 - PubMed Abstract:
T-cell priming occurs when a naïve T cell recognizes cognate peptide-MHC complexes on an activated antigen-presenting cell. The circumstances of this initial priming have ramifications on the fate of the newly primed T cell. Newly primed CD8 + T cells can embark onto different trajectories, with some becoming short-lived effector cells and others adopting a tissue resident or memory cell fate. To determine whether T-cell priming influences the quality of the effector T-cell response to tumors, we used transnuclear CD8 + T cells that recognize the melanoma antigen TRP1 using TRP1 high or TRP1 low TCRs that differ in both affinity and fine specificity. From a series of altered peptide ligands, we identified a point mutation (K8) in a nonanchor residue that, when analyzed crystallographically and biophysically, destabilized the peptide interaction with the MHC binding groove. In vitro , the K8 peptide induced robust proliferation of both TRP1 high and TRP1 low CD8 + T cells but did not induce expression of PD-1. Cytokine production from K8-stimulated TRP1 cells was minimal, whereas cytotoxicity was increased. Upon transfer into B16 tumor-bearing mice, the reference peptide (TRP1-M9)- and K8-stimulated TRP1 cells were equally effective at controlling tumor growth but accomplished this through different mechanisms. TRP1-M9-stimulated cells produced more IFNγ, whereas K8-stimulated cells accumulated to higher numbers and were more cytotoxic. We, therefore, conclude that TCR recognition of weakly binding peptides during priming can skew the effector function of tumor-specific CD8 + T cells.
- Department of Cancer Immunology and Virology, Dana-Farber Cancer Institute, Boston, Massachusetts.
Organizational Affiliation: