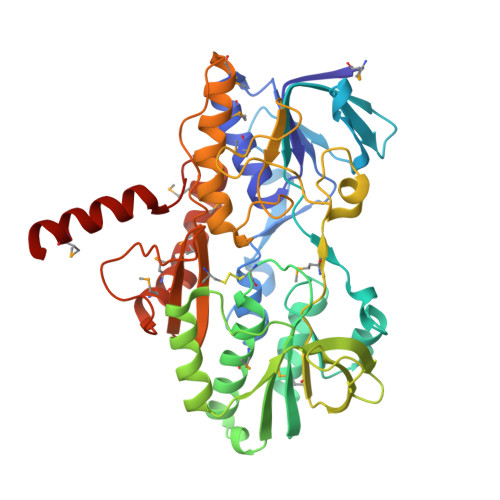
X-Ray Structure of Human Sulfide:Quinone Oxidoreductase: Insights into the Mechanism of Mitochondrial Hydrogen Sulfide Oxidation.
Jackson, M.R., Loll, P.J., Jorns, M.S.(2019) Structure 27: 794-805.e4
- PubMed: 30905673 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.str.2019.03.002
- Primary Citation Related Structures:
6MO6, 6MP5 - PubMed Abstract:
Hydrogen sulfide (H 2 S) is a gasotransmitter exhibiting pivotal functions in diverse biological processes, including activation of multiple cardioprotective pathways. Sulfide:quinone oxidoreductase (SQOR) is an integral membrane flavoprotein that catalyzes the first step in the mitochondrial metabolism of H 2 S. As such, it plays a critical role in controlling physiological levels of the gasotransmitter and has attracted keen interest as a potential drug target. We report the crystal structure of human SQOR, unraveling the molecular basis for the enzyme's ability to catalyze sulfane sulfur transfer reactions with structurally diverse acceptors. We demonstrate that human SQOR contains unique features: an electropositive surface depression implicated as a binding site for sulfane sulfur acceptors and postulated to funnel negatively charged substrates to a hydrophilic H 2 S-oxidizing active site, which is connected to a hydrophobic internal tunnel that binds coenzyme Q. These findings support a proposed model for catalysis and open the door for structure-based drug design.
- Biochemistry and Molecular Biology, Drexel University College of Medicine, 245 N. 15(th) Street, Philadelphia, PA 19102, USA.
Organizational Affiliation: