cAMP-dependent protein kinase A in complex with RyR2 peptide (2799-2810)
Haji-Ghassemi, O., Yuchi, Z., van Petegem, F.(2019) Mol Cell
Experimental Data Snapshot
Starting Model: experimental
View more details
(2019) Mol Cell
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| cAMP-dependent protein kinase catalytic subunit alpha | A [auth C], B [auth E] | 339 | Mus musculus | Mutation(s): 0 Gene Names: Prkaca, Pkaca EC: 2.7.11.11 | 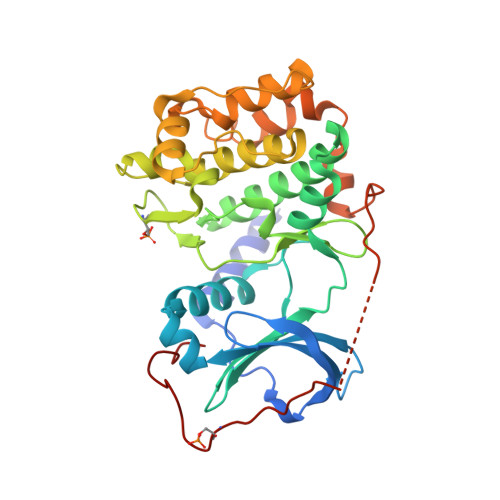 |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P05132 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Ryanodine receptor 2 | C [auth F], D | 209 | Mus musculus | Mutation(s): 1 Gene Names: Ryr2 | 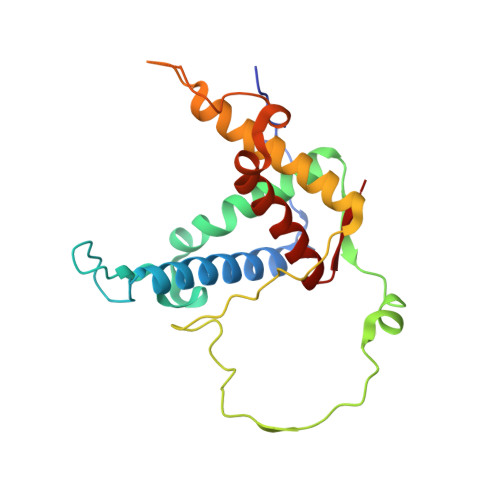 |
UniProt & NIH Common Fund Data Resources | |||||
IMPC: MGI:99685 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | E9Q401 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 5 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| ANP (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | F [auth C], I [auth E] | PHOSPHOAMINOPHOSPHONIC ACID-ADENYLATE ESTER C10 H17 N6 O12 P3 PVKSNHVPLWYQGJ-KQYNXXCUSA-N |  | ||
| PGE Download:Ideal Coordinates CCD File | G [auth C] | TRIETHYLENE GLYCOL C6 H14 O4 ZIBGPFATKBEMQZ-UHFFFAOYSA-N |  | ||
| EDO Download:Ideal Coordinates CCD File | J [auth E] | 1,2-ETHANEDIOL C2 H6 O2 LYCAIKOWRPUZTN-UHFFFAOYSA-N |  | ||
| ACT Download:Ideal Coordinates CCD File | E [auth C] | ACETATE ION C2 H3 O2 QTBSBXVTEAMEQO-UHFFFAOYSA-M |  | ||
| CL Download:Ideal Coordinates CCD File | H [auth C], K [auth E], L [auth F] | CHLORIDE ION Cl VEXZGXHMUGYJMC-UHFFFAOYSA-M |  | ||
| Modified Residues 2 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Type | Formula | 2D Diagram | Parent |
| SEP Query on SEP | A [auth C], B [auth E] | L-PEPTIDE LINKING | C3 H8 N O6 P |  | SER |
| TPO Query on TPO | A [auth C], B [auth E] | L-PEPTIDE LINKING | C4 H10 N O6 P |  | THR |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 58.451 | α = 105.51 |
| b = 74.944 | β = 90.14 |
| c = 88.69 | γ = 94.39 |
| Software Name | Purpose |
|---|---|
| REFMAC | refinement |
| HKL-2000 | data scaling |
| PDB_EXTRACT | data extraction |
| HKL-2000 | data reduction |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Canadian Institutes of Health Research (CIHR) | Canada | PJT 153305 |