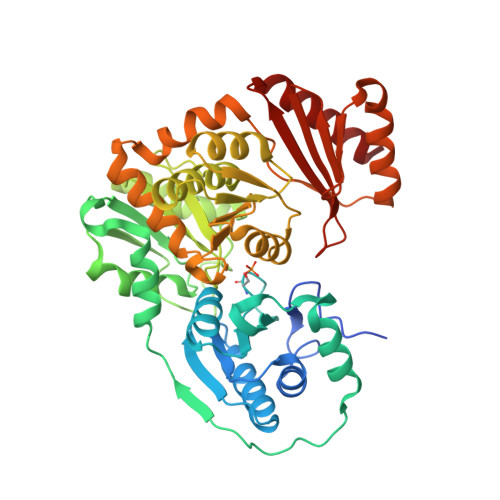
Inhibitory Evaluation of alpha PMM/PGM fromPseudomonas aeruginosa: Chemical Synthesis, Enzyme Kinetics, and Protein Crystallographic Study.
Zhu, J.S., Stiers, K.M., Soleimani, E., Groves, B.R., Beamer, L.J., Jakeman, D.L.(2019) J Org Chem 84: 9627-9636
- PubMed: 31264865 Search on PubMed
- DOI: https://doi.org/10.1021/acs.joc.9b01305
- Primary Citation Related Structures:
6MLF, 6MLH, 6MLW, 6MNV - PubMed Abstract:
α-Phosphomannomutase/phosphoglucomutase (αPMM/PGM) from P. aeruginosa is involved in bacterial cell wall assembly and is implicated in P. aeruginosa virulence, yet few studies have addressed αPMM/PGM inhibition from this important Gram-negative bacterial human pathogen. Four structurally different α-d-glucopyranose 1-phosphate (αG1P) derivatives including 1- C -fluoromethylated analogues ( 1 - 3 ), 1,2-cyclic phosph(on)ate analogues ( 4 - 6 ), isosteric methylene phosphono analogues ( 7 and 8 ), and 6-fluoro-αG1P ( 9 ), were synthesized and assessed as potential time-dependent or reversible αPMM/PGM inhibitors. The resulting kinetic data were consistent with the crystallographic structures of the highly homologous Xanthomonas citri αPGM with inhibitors 3 and 7 - 9 binding to the enzyme active site (1.65-1.9 Å). These structural and kinetic insights will enhance the design of future αPMM/PGM inhibitors.
- College of Pharmacy , Dalhousie University , 5968 College Street , Halifax , Nova Scotia B3H 4R2 , Canada.
Organizational Affiliation: