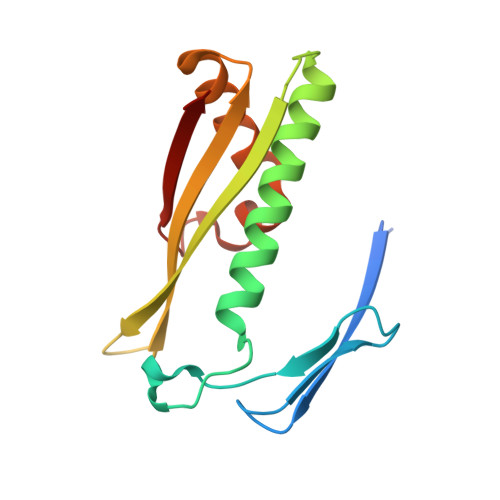
Crystal structure of an organic hydroperoxide resistance protein OsmC, predicted redox protein, regulator of sulfide bond formation from Legionella pneumophila
Edwards, T.E., Delker, S.L., Horanyi, P.S., Lorimer, D.D., Seattle Structural Genomics Center for Infectious Disease (SSGCID)To be published.