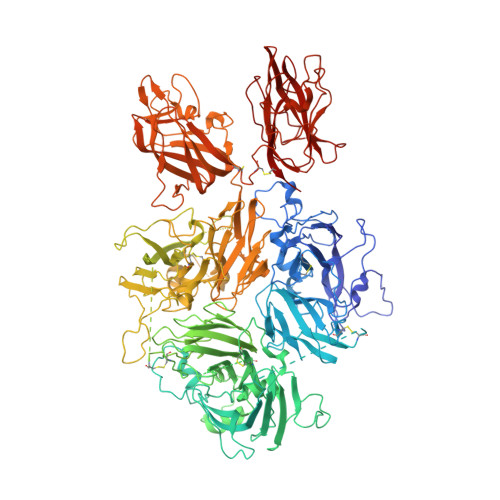
The 3.2 angstrom structure of a bioengineered variant of blood coagulation factor VIII indicates two conformations of the C2 domain.
Smith, I.W., d'Aquino, A.E., Coyle, C.W., Fedanov, A., Parker, E.T., Denning, G., Spencer, H.T., Lollar, P., Doering, C.B., Spiegel Jr., P.C.(2020) J Thromb Haemost 18: 57-69
- PubMed: 31454152 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1111/jth.14621
- Primary Citation Related Structures:
6MF0, 6MF2 - PubMed Abstract:
Coagulation factor VIII represents one of the oldest protein-based therapeutics, serving as an effective hemophilia A treatment for half a century. Optimal treatment consists of repeated intravenous infusions of blood coagulation factor VIII (FVIII) per week for life. Despite overall treatment success, significant limitations remain, including treatment invasiveness, duration, immunogenicity, and cost. These issues have inspired research into the development of bioengineered FVIII products and gene therapies. To structurally characterize a bioengineered construct of FVIII, termed ET3i, which is a human/porcine chimeric B domain-deleted heterodimer with improved expression and slower A2 domain dissociation following proteolytic activation by thrombin. The structure of ET3i was characterized with X-ray crystallography and tandem mass spectrometry-based glycoproteomics. Here, we report the 3.2 Å crystal structure of ET3i and characterize the distribution of N-linked glycans with LC-MS/MS glycoproteomics. This structure shows remarkable conservation with the human FVIII protein and provides a detailed view of the interface between the A2 domain and the remaining FVIII structure. With two FVIII molecules in the crystal, we observe two conformations of the C2 domain relative to the remaining FVIII structure. The improved model and stereochemistry of ET3i served as a scaffold to generate an improved, refined structure of human FVIII. With the original datasets at 3.7 Å and 4.0 Å resolution, this new structure resulted in improved refinement statistics. These improved structures yield a more confident model for next-generation engineering efforts to develop FVIII therapeutics with longer half-lives, higher expression levels, and lower immunogenicity.
- Department of Chemistry, Western Washington University, Bellingham, Washington.
Organizational Affiliation: