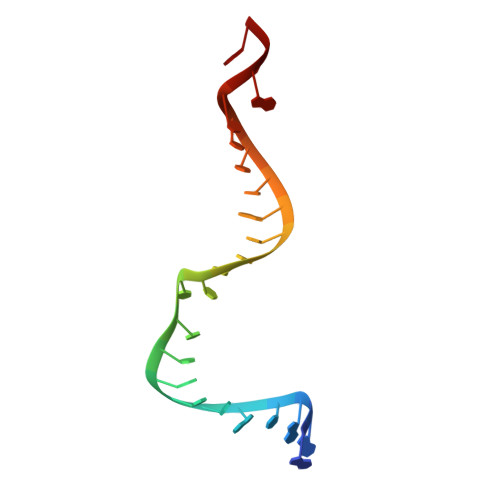
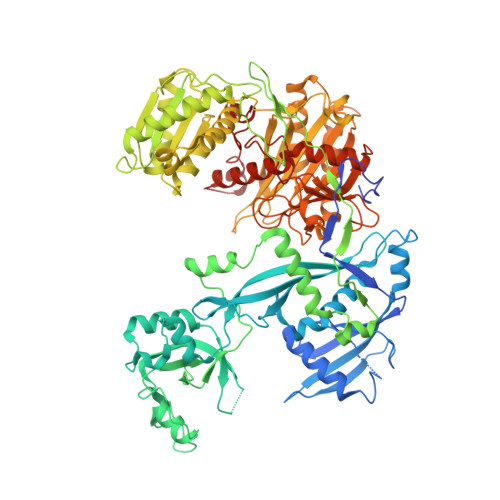
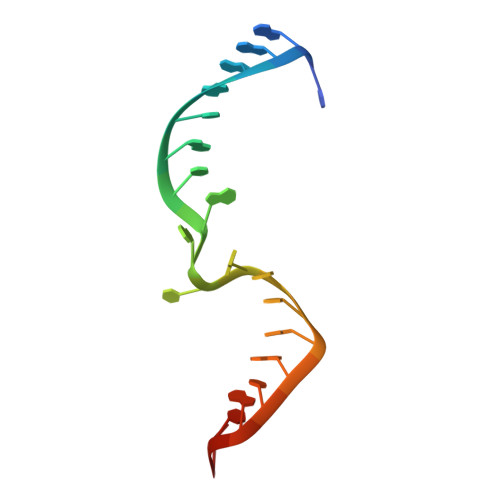
Structural Basis for Target-Directed MicroRNA Degradation.
Sheu-Gruttadauria, J., Pawlica, P., Klum, S.M., Wang, S., Yario, T.A., Schirle Oakdale, N.T., Steitz, J.A., MacRae, I.J.(2019) Mol Cell 75: 1243-1255.e7
- PubMed: 31353209 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.molcel.2019.06.019
- Primary Citation Related Structures:
6MDZ, 6MFN, 6MFR, 6NIT - PubMed Abstract:
MicroRNAs (miRNAs) broadly regulate gene expression through association with Argonaute (Ago), which also protects miRNAs from degradation. However, miRNA stability is known to vary and is regulated by poorly understood mechanisms. A major emerging process, termed target-directed miRNA degradation (TDMD), employs specialized target RNAs to selectively bind to miRNAs and induce their decay. Here, we report structures of human Ago2 (hAgo2) bound to miRNAs and TDMD-inducing targets. miRNA and target form a bipartite duplex with an unpaired flexible linker. hAgo2 cannot physically accommodate the RNA, causing the duplex to bend at the linker and display the miRNA 3' end for enzymatic attack. Altering 3' end display by changing linker flexibility, changing 3' end complementarity, or mutationally inducing 3' end release impacts TDMD efficiency, leading to production of distinct 3'-miRNA isoforms in cells. Our results uncover the mechanism driving TDMD and reveal 3' end display as a key determinant regulating miRNA activity via 3' remodeling and/or degradation.
- Department of Integrative Structural and Computational Biology, The Scripps Research Institute, La Jolla, CA 92037, USA.
Organizational Affiliation: