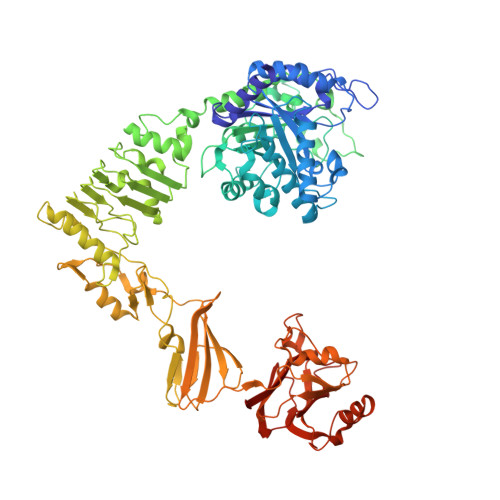
Molecular Basis of Broad SpectrumN-Glycan Specificity and Processing of Therapeutic IgG Monoclonal Antibodies by Endoglycosidase S2.
Klontz, E.H., Trastoy, B., Deredge, D., Fields, J.K., Li, C., Orwenyo, J., Marina, A., Beadenkopf, R., Gunther, S., Flores, J., Wintrode, P.L., Wang, L.X., Guerin, M.E., Sundberg, E.J.(2019) ACS Cent Sci 5: 524-538
- PubMed: 30937380 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acscentsci.8b00917
- Primary Citation Related Structures:
6E58, 6MDS, 6MDV - PubMed Abstract:
Immunoglobulin G (IgG) glycosylation critically modulates antibody effector functions. Streptococcus pyogenes secretes a unique endo-β- N -acetylglucosaminidase, EndoS2, which deglycosylates the conserved N -linked glycan at Asn297 on IgG Fc to eliminate its effector functions and evade the immune system. EndoS2 and specific point mutants have been used to chemoenzymatically synthesize antibodies with customizable glycosylation for gain of functions. EndoS2 is useful in these schemes because it accommodates a broad range of N -glycans, including high-mannose, complex, and hybrid types; however, its mechanism of substrate recognition is poorly understood. We present crystal structures of EndoS2 alone and bound to complex and high-mannose glycans; the broad N -glycan specificity is governed by critical loops that shape the binding site of EndoS2. Furthermore, hydrolytic experiments, domain-swap chimeras, and hydrogen-deuterium exchange mass spectrometry reveal the importance of the carbohydrate-binding module in the mechanism of IgG recognition by EndoS2, providing insights into engineering enzymes to catalyze customizable glycosylation reactions.
- Institute of Human Virology, Department of Microbiology & Immunology, and Program in Molecular Microbiology & Immunology, University of Maryland School of Medicine, Baltimore, Maryland 21201, United States.
Organizational Affiliation: