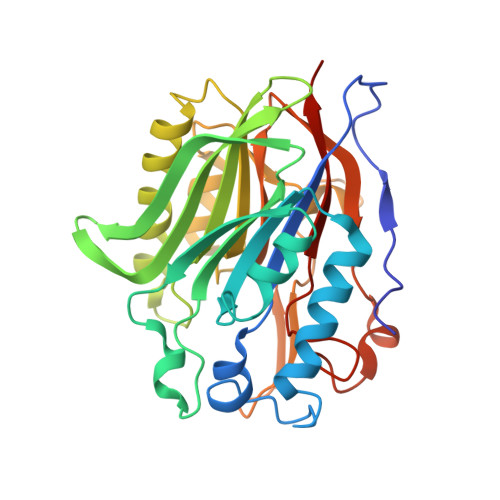
Crystal Structure of Human Nocturnin Catalytic Domain.
Estrella, M.A., Du, J., Korennykh, A.(2018) Sci Rep 8: 16294-16294
- PubMed: 30389976 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-018-34615-0
- Primary Citation Related Structures:
6MAL - PubMed Abstract:
Nocturnin (NOCT) helps the circadian clock to adjust metabolism according to day and night activity. NOCT is upregulated in early evening and it has been proposed that NOCT serves as a deadenylase for metabolic enzyme mRNAs. We present a 2.7-Å crystal structure of the catalytic domain of human NOCT. Our structure shows that NOCT has a close overall similarity to CCR4 deadenylase family members, PDE12 and CNOT6L, and to a DNA repair enzyme TDP2. All the key catalytic residues present in PDE12, CNOT6L and TDP2 are conserved in NOCT and have the same conformations. However, we observe substantial differences in the surface properties of NOCT, an unexpectedly narrow active site pocket, and conserved structural elements in the vicinity of the catalytic center, which are unique to NOCT and absent in the deadenylases PDE12/CNOT6L. Moreover, we show that in contrast to human PDE12 and CNOT6L, NOCT is completely inactive against poly-A RNA. Our work thus reveals the structure of an intriguing circadian protein and suggests that NOCT has considerable differences from the related deadenylases, which may point to a unique cellular function of this enzyme.
- Department of Molecular Biology, Princeton, NJ, 08544, USA.
Organizational Affiliation: