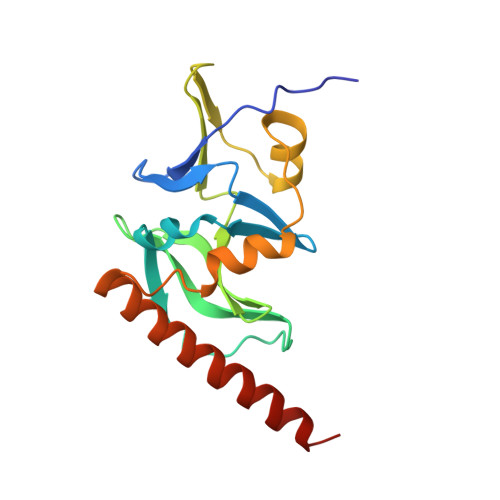
Insights into the structure and RNA-binding specificity of Caenorhabditis elegans Dicer-related helicase 3 (DRH-3).
Li, K., Zheng, J., Wirawan, M., Trinh, N.M., Fedorova, O., Griffin, P.R., Pyle, A.M., Luo, D.(2021) Nucleic Acids Res 49: 9978-9991
- PubMed: 34403472 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkab712
- Primary Citation Related Structures:
6M6Q, 6M6R, 6M6S - PubMed Abstract:
DRH-3 is critically involved in germline development and RNA interference (RNAi) facilitated chromosome segregation via the 22G-siRNA pathway in Caenorhabditis elegans. DRH-3 has similar domain architecture to RIG-I-like receptors (RLRs) and belongs to the RIG-I-like RNA helicase family. The molecular understanding of DRH-3 and its function in endogenous RNAi pathways remains elusive. In this study, we solved the crystal structures of the DRH-3 N-terminal domain (NTD) and the C-terminal domains (CTDs) in complex with 5'-triphosphorylated RNAs. The NTD of DRH-3 adopts a distinct fold of tandem caspase activation and recruitment domains (CARDs) structurally similar to the CARDs of RIG-I and MDA5, suggesting a signaling function in the endogenous RNAi biogenesis. The CTD preferentially recognizes 5'-triphosphorylated double-stranded RNAs bearing the typical features of secondary siRNA transcripts. The full-length DRH-3 displays unique structural dynamics upon binding to RNA duplexes that differ from RIG-I or MDA5. These features of DRH-3 showcase the evolutionary divergence of the Dicer and RLR family of helicases.
- Lee Kong Chian School of Medicine, Nanyang Technological University, EMB 03-07, 59 Nanyang Drive 636921, Singapore.
Organizational Affiliation: