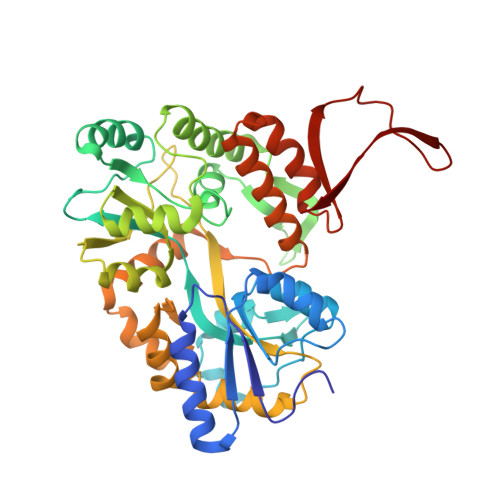
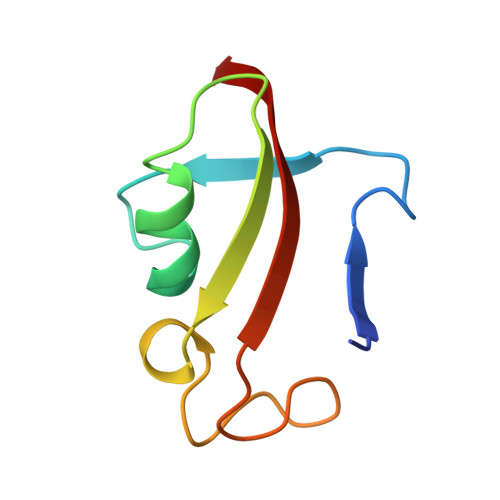
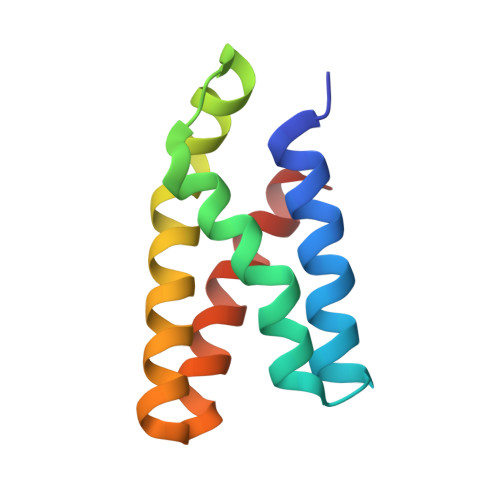
Rational design and implementation of a chemically inducible heterotrimerization system.
Wu, H.D., Kikuchi, M., Dagliyan, O., Aragaki, A.K., Nakamura, H., Dokholyan, N.V., Umehara, T., Inoue, T.(2020) Nat Methods 17: 928-936
- PubMed: 32747768 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41592-020-0913-x
- Primary Citation Related Structures:
6M4U, 6M4V, 6M4W - PubMed Abstract:
Chemically inducible dimerization (CID) uses a small molecule to induce binding of two different proteins. CID tools such as the FK506-binding protein-FKBP-rapamycin-binding- (FKBP-FRB)-rapamycin system have been widely used to probe molecular events inside and outside cells. While various CID tools are available, chemically inducible trimerization (CIT) does not exist, due to inherent challenges in designing a chemical that simultaneously binds three proteins with high affinity and specificity. Here, we developed CIT by rationally splitting FRB and FKBP. Cellular and structural datasets showed efficient trimerization of split pairs of FRB or FKBP with full-length FKBP or FRB, respectively, by rapamycin. CIT rapidly induced tri-organellar junctions and perturbed intended membrane lipids exclusively at select membrane contact sites. By conferring one additional condition to what is achievable with CID, CIT expands the types of manipulation in single live cells to address cell biology questions otherwise intractable and engineer cell functions for future synthetic biology applications.
- Department of Biomedical Engineering, Johns Hopkins University School of Medicine, Baltimore, MD, USA.
Organizational Affiliation: