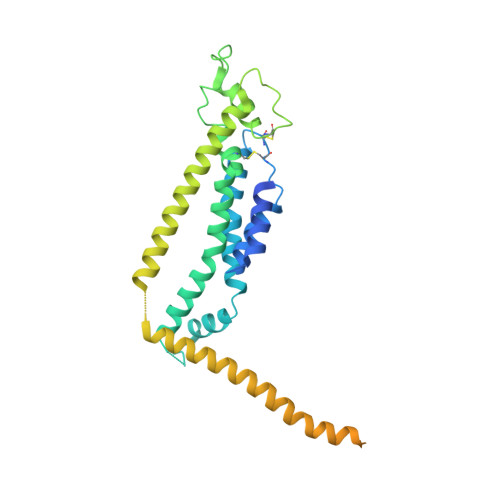
Cryo-EM structure of the calcium homeostasis modulator 1 channel.
Ren, Y., Wen, T., Xi, Z., Li, S., Lu, J., Zhang, X., Yang, X., Shen, Y.(2020) Sci Adv 6: eaba8161-eaba8161
- PubMed: 32832630 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.aba8161
- Primary Citation Related Structures:
6LYG - PubMed Abstract:
Calcium homeostasis modulator 1 (CALHM1) is a voltage-gated ATP release channel that plays an important role in neural gustatory signaling and the pathogenesis of Alzheimer's disease. Here, we present a cryo-electron microscopy structure of full-length Ca 2+ -free CALHM1 from Danio rerio at an overall resolution of 3.1 Å. Our structure reveals an octameric architecture with a wide pore diameter of ~20 Å, presumably representing the active conformation. The overall structure is substantially different from that of the isoform CALHM2, which forms both undecameric hemichannels and gap junctions. The N-terminal small helix folds back to the pore and forms an antiparallel interaction with transmembrane helix 1. Structural analysis revealed that the extracellular loop 1 region within the dimer interface may contribute to oligomeric assembly. A positive potential belt inside the pore was identified that may modulate ion permeation. Our structure offers insights into the assembly and gating mechanism of the CALHM1 channel.
- State Key Laboratory of Medicinal Chemical Biology and College of Pharmacy, Nankai University, Tianjin 300350, China.
Organizational Affiliation: