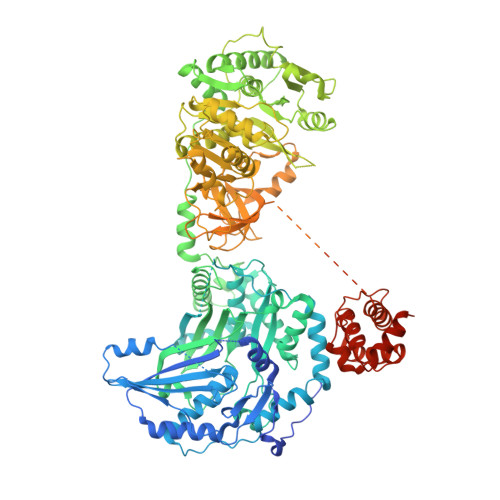
Structural and Functional Analyses of the Tridomain-Nonribosomal Peptide Synthetase FmoA3 for 4-Methyloxazoline Ring Formation.
Katsuyama, Y., Sone, K., Harada, A., Kawai, S., Urano, N., Adachi, N., Moriya, T., Kawasaki, M., Shin-Ya, K., Senda, T., Ohnishi, Y.(2021) Angew Chem Int Ed Engl 60: 14554-14562
- PubMed: 33783097
- DOI: https://doi.org/10.1002/anie.202102760
- Primary Citation of Related Structures:
6LTA, 6LTB, 6LTC, 6LTD - PubMed Abstract:
Nonribosomal peptide synthetases (NRPSs) are attractive targets for bioengineering to generate useful peptides. FmoA3 is a single modular NRPS composed of heterocyclization (Cy), adenylation (A), and peptidyl carrier protein (PCP) domains. It uses α-methyl-l-serine to synthesize a 4-methyloxazoline ring, probably with another Cy domain in the preceding module FmoA2. Here, we determined the head-to-tail homodimeric structures of FmoA3 by X-ray crystallography (apo-form, with adenylyl-imidodiphosphate and α-methyl-l-seryl-AMP) and cryogenic electron microscopy single particle analysis, and performed site-directed mutagenesis experiments. The data revealed that α-methyl-l-serine can be accommodated in the active site because of the extra space around Ala688. The Cy domains of FmoA2 and FmoA3 catalyze peptide bond formation and heterocyclization, respectively. FmoA3's Cy domain seems to lose its donor PCP binding activity. The collective data support a proposed catalytic cycle of FmoA3.
- Department of Biotechnology, Graduate School of Agricultural and Life science, The University of Tokyo, 1-1-1 Yayoi, Bunkyo-ku, Tokyo, 113-8657, Japan.
Organizational Affiliation: