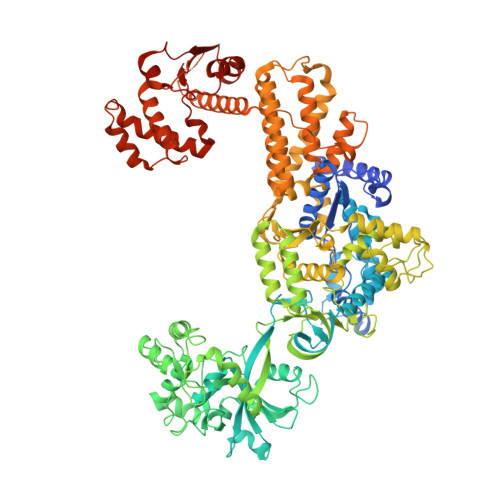
Molecular basis of the multifaceted functions of human leucyl-tRNA synthetase in protein synthesis and beyond.
Liu, R.J., Long, T., Li, H., Zhao, J., Li, J., Wang, M., Palencia, A., Lin, J., Cusack, S., Wang, E.D.(2020) Nucleic Acids Res 48: 4946-4959
- PubMed: 32232361 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkaa189
- Primary Citation Related Structures:
6LPF, 6LR6 - PubMed Abstract:
Human cytosolic leucyl-tRNA synthetase (hcLRS) is an essential and multifunctional enzyme. Its canonical function is to catalyze the covalent ligation of leucine to tRNALeu, and it may also hydrolyze mischarged tRNAs through an editing mechanism. Together with eight other aminoacyl-tRNA synthetases (AaRSs) and three auxiliary proteins, it forms a large multi-synthetase complex (MSC). Beyond its role in translation, hcLRS has an important moonlight function as a leucine sensor in the rapamycin complex 1 (mTORC1) pathway. Since this pathway is active in cancer development, hcLRS is a potential target for anti-tumor drug development. Moreover, LRS from pathogenic microbes are proven drug targets for developing antibiotics, which however should not inhibit hcLRS. Here we present the crystal structure of hcLRS at a 2.5 Å resolution, the first complete structure of a eukaryotic LRS, and analyze the binding of various compounds that target different sites of hcLRS. We also deduce the assembly mechanism of hcLRS into the MSC through reconstitution of the entire mega complex in vitro. Overall, our study provides the molecular basis for understanding both the multifaceted functions of hcLRS and for drug development targeting these functions.
- School of Life Science and Technology, ShanghaiTech University, 100 Haike Road, Shanghai 201210, P.R. China.
Organizational Affiliation: