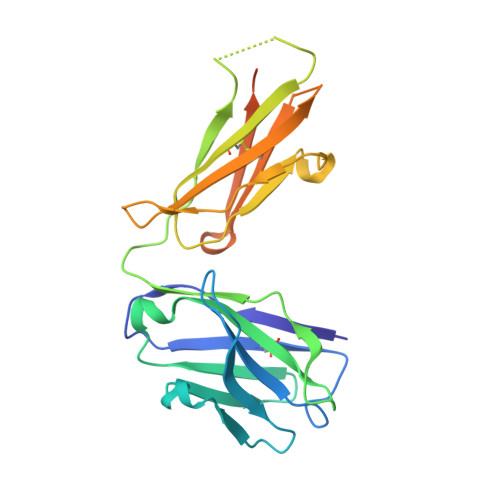
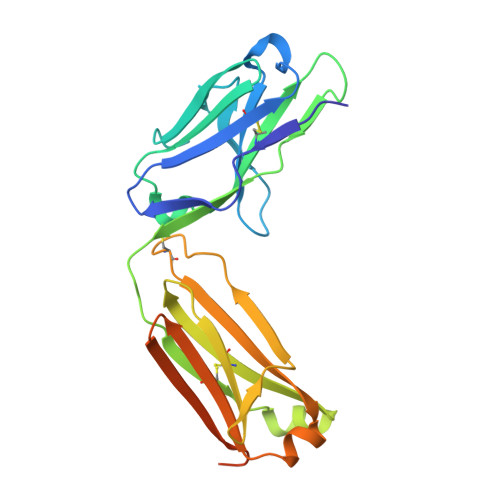
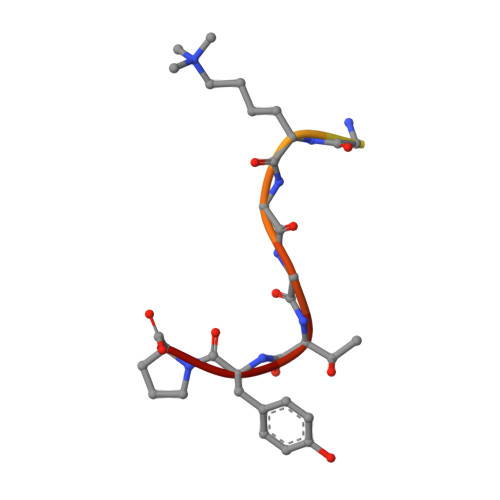
Structural basis for antigen recognition by methylated lysine-specific antibodies.
Ishii, M., Nakakido, M., Caaveiro, J.M.M., Kuroda, D., Okumura, C.J., Maruyama, T., Entzminger, K., Tsumoto, K.(2020) J Biological Chem 296: 100176-100176
- PubMed: 33303630 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.RA120.015996
- Primary Citation Related Structures:
6LDV, 6LDW, 6LDX, 6LDY - PubMed Abstract:
Proteins are modulated by a variety of posttranslational modifications including methylation. Despite its importance, the majority of protein methylation modifications discovered by mass spectrometric analyses are functionally uncharacterized, partly owing to the difficulty in obtaining reliable methylsite-specific antibodies. To elucidate how functional methylsite-specific antibodies recognize the antigens and lead to the development of a novel method to create such antibodies, we use an immunized library paired with phage display to create rabbit monoclonal antibodies recognizing trimethylated Lys260 of MAP3K2 as a representative substrate. We isolated several methylsite-specific antibodies that contained unique complementarity determining region sequence. We characterized the mode of antigen recognition by each of these antibodies using structural and biophysical analyses, revealing the molecular details, such as binding affinity toward methylated/nonmethylated antigens and structural motif that is responsible for recognition of the methylated lysine residue, by which each antibody recognized the target antigen. In addition, the comparison with the results of Western blotting analysis suggests a critical antigen recognition mode to generate cross-reactivity to protein and peptide antigen of the antibodies. Computational simulations effectively recapitulated our biophysical data, capturing the antibodies of differing affinity and specificity. Our exhaustive characterization provides molecular architectures of functional methylsite-specific antibodies and thus should contribute to the development of a general method to generate functional methylsite-specific antibodies by de novo design.
- Department of Bioengineering, School of Engineering, The University of Tokyo, Tokyo, Japan.
Organizational Affiliation: