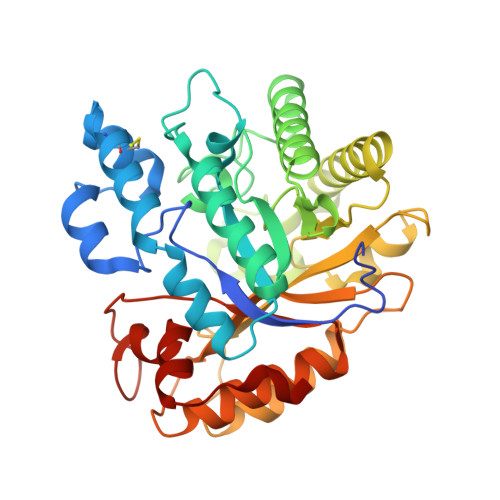
Aspergillus oryzae Rutinosidase: Biochemical and Structural Investigation.
Makabe, K., Hirota, R., Shiono, Y., Tanaka, Y., Koseki, T.(2021) Appl Environ Microbiol 87
- PubMed: 33218993 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/AEM.02438-20
- Primary Citation Related Structures:
6LA0 - PubMed Abstract:
The rutinosidase (Rut)-encoding gene Aorut has been expressed in Pichia pastoris with its native signal sequence from Aspergillus oryzae Biochemical and structural investigation of the purified recombinant mature A. oryzae Rut ( Ao Rut), designated r Ao RutM, was performed in this study. A 1.7-Å resolution crystal structure of r Ao RutM was determined, which is an essential step forward in the utilization of Ao Rut as a potential catalyst. The crystal structure of r Ao RutM was represented by a (β/α) 8 TIM barrel fold with structural similarity to that of rutinosidase from Aspergillus niger ( An Rut) and an exo-β-(1,3)-glucanase from Candida albicans The crystal structure revealed that the catalytic site was located in a deep cleft, similarly to An Rut, and that internal cavities and water molecules were also present. Purified r Ao RutM hydrolyzed not only 7- O -linked and 3- O -linked flavonoid rutinosides but also 7- O -linked and 3- O -linked flavonoid glucosides. r Ao RutM displayed high catalytic activity toward quercetin 3- O -linked substrates such as rutin and isoquercitrin, rather than to the 7- O -linked substrate, quercetin-7- O -glucoside. Unexpectedly, purified r Ao RutM exhibited increased thermostability after treatment with endo-β- N -acetylglucosaminidase H. Circular dichroism (CD) spectra of purified intact r Ao RutM and of the enzyme after N -deglycosylation showed a typical α-helical CD profile; however, the molar ellipticity values of the peaks at 208 nm and 212 nm differed. The K m and k cat values for the substrates modified by rutinose were higher than those for the substrates modified by β-d-glucose. IMPORTANCE Flavonoid glycosides constitute a class of secondary metabolites widely distributed in nature. These compounds are involved in bitter taste or clouding in plant-based foods or beverages, respectively. Flavonoid glycoside degradation can proceed through two alternative enzymatic pathways: one that is mediated by monoglycosidases and another that is catalyzed by a diglycosidase. The present report on the biochemical and structural investigation of A. oryzae rutinosidase provides a potential biocatalyst for industrial applications of flavonoids.
- Graduate School of Science and Engineering, Faculty of Engineering, Yamagata University, Yonezawa, Japan makabe@yz.yamagata-u.ac.jp tkoseki@tds1.tr.yamagata-u.ac.jp.
Organizational Affiliation: