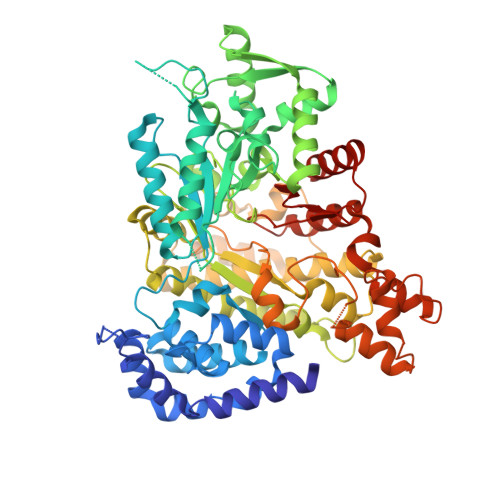
TAS1553, a novel small molecule ribonucleotide reductase (RNR) subunit interaction inhibitor, displays remarkable anti-tumor activity
Miyahara, S., Hara, S., Chong, K.T., Suzuki, T., Ogino, Y., Hoshino, T., Tsukioka, S., Yano, W., Suzuki, M., Otsu, Y., Yonekura, T., Ito, S., Terasaka, M., Suzuki, T., Hashimoto, A.To be published.