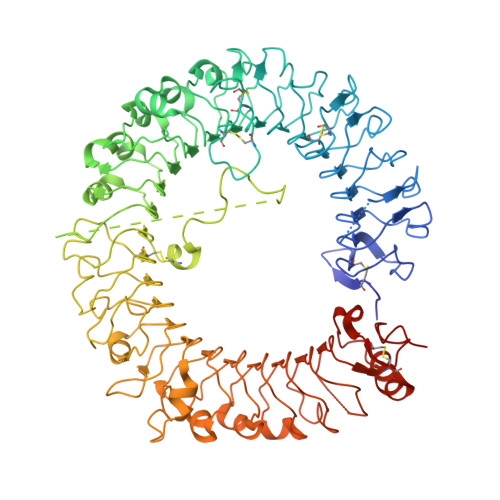
Rationally Designed Small-Molecule Inhibitors Targeting an Unconventional Pocket on the TLR8 Protein-Protein Interface.
Jiang, S., Tanji, H., Yin, K., Zhang, S., Sakaniwa, K., Huang, J., Yang, Y., Li, J., Ohto, U., Shimizu, T., Yin, H.(2020) J Med Chem 63: 4117-4132
- PubMed: 32233366 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.9b02128
- Primary Citation Related Structures:
6KYA - PubMed Abstract:
Rational designs of small-molecule inhibitors targeting protein-protein interfaces have met little success. Herein, we have designed a series of triazole derivatives with a novel scaffold to specifically intervene with the interaction of TLR8 homomerization. In multiple assays, TH1027 was identified as a highly potent and specific inhibitor of TLR8. A successful solution of the X-ray crystal structure of TLR8 in complex with TH1027 provided an in-depth mechanistic insight into its binding mode, validating that TH1027 was located between two TLR8 monomers and recognized as an unconventional pocket, thereby preventing TLR8 from activation. Further biological evaluations showed that TH1027 dose-dependently suppressed the TLR8-mediated inflammatory responses in both human monocyte cell lines, peripheral blood mononuclear cells, and rheumatoid arthritis patient specimens, suggesting a strong therapeutic potential against autoimmune diseases.
- Graduate School of Pharmaceutical Sciences, The University of Tokyo, Tokyo 113-0033, Japan.
Organizational Affiliation: