Crystal structure of human alpha2C adrenergic G protein-coupled receptor.
Chen, X.Y., Wu, D., Wu, L.J., Han, G.W., Guo, Y., Zhong, G.S.To be published.
Experimental Data Snapshot
Starting Model: experimental
View more details
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Alpha-2C adrenergic receptor | 496 | Homo sapiens | Mutation(s): 0 Membrane Entity: Yes | 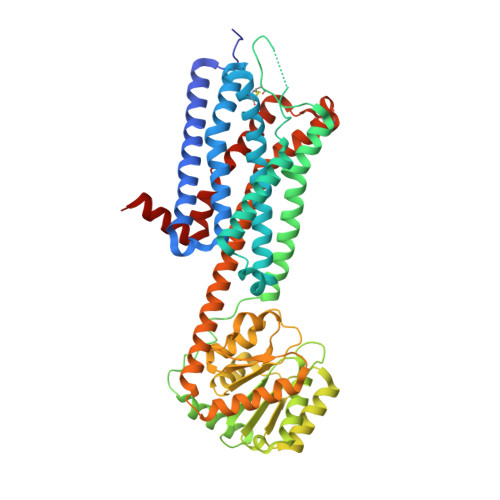 | |
UniProt & NIH Common Fund Data Resources | |||||
PHAROS: P18825 GTEx: ENSG00000184160 | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Groups | Q9V2J8P18825 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 4 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| CLR Download:Ideal Coordinates CCD File | D [auth A], M [auth B] | CHOLESTEROL C27 H46 O HVYWMOMLDIMFJA-DPAQBDIFSA-N |  | ||
| E33 Download:Ideal Coordinates CCD File | C [auth A], L [auth B] | (8~{a}~{R},12~{a}~{S},13~{a}~{R})-12-ethylsulfonyl-3-methoxy-5,6,8,8~{a},9,10,11,12~{a},13,13~{a}-decahydroisoquinolino[2,1-g][1,6]naphthyridine C19 H28 N2 O3 S UMGBFFAJXFXOIL-AYOQOUSVSA-N |  | ||
| OLC Download:Ideal Coordinates CCD File | E [auth A] F [auth A] G [auth A] H [auth A] I [auth A] | (2R)-2,3-dihydroxypropyl (9Z)-octadec-9-enoate C21 H40 O4 RZRNAYUHWVFMIP-GDCKJWNLSA-N |  | ||
| OLA Download:Ideal Coordinates CCD File | J [auth A], K [auth A] | OLEIC ACID C18 H34 O2 ZQPPMHVWECSIRJ-KTKRTIGZSA-N |  | ||
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 74.48 | α = 90 |
| b = 78.74 | β = 90 |
| c = 190.45 | γ = 90 |
| Software Name | Purpose |
|---|---|
| BUSTER | refinement |
| XDS | data reduction |
| XSCALE | data scaling |
| PHASER | phasing |
| Funding Organization | Location | Grant Number |
|---|---|---|
| National Natural Science Foundation of China | China | 31771130 |