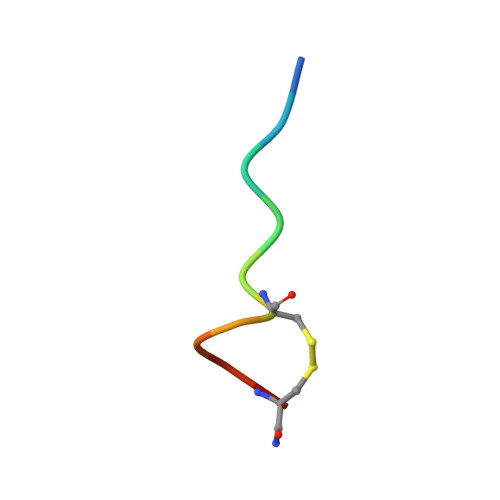
Structure and allosteric activity of a single-disulfide conopeptide fromConus zonatusat human alpha 3 beta 4 and alpha 7 nicotinic acetylcholine receptors.
Mohan, M.K., Abraham, N., R P, R., Jayaseelan, B.F., Ragnarsson, L., Lewis, R.J., Sarma, S.P.(2020) J Biological Chem 295: 7096-7112
- PubMed: 32234761 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.RA119.012098
- Primary Citation Related Structures:
6KMY, 6KN2, 6KN3, 6KNO, 6KNP - PubMed Abstract:
Conopeptides are neurotoxic peptides in the venom of marine cone snails and have broad therapeutic potential for managing pain and other conditions. Here, we identified the single-disulfide peptides Czon1107 and Cca1669 from the venoms of Conus zonatus and Conus caracteristicus , respectively. We observed that Czon1107 strongly inhibits the human α3β4 (IC 50 15.7 ± 3.0 μm) and α7 (IC 50 77.1 ± 0.05 μm) nicotinic acetylcholine receptor (nAChR) subtypes, but the activity of Cca1669 remains to be identified. Czon1107 acted at a site distinct from the orthosteric receptor site. Solution NMR experiments revealed that Czon1107 exists in equilibrium between conformational states that are the result of a key Ser 4 -Pro 5 cis-trans isomerization. Moreover, we found that the X-Pro amide bonds in the inter-cysteine loop are rigidly constrained to cis conformations. Structure-activity experiments of Czon1107 and its variants at positions P5 and P7 revealed that the conformation around the X-Pro bonds ( cis-trans ) plays an important role in receptor subtype selectivity. The cis conformation at the Cys 6 -Pro 7 peptide bond was essential for α3β4 nAChR subtype allosteric selectivity. In summary, we have identified a unique single-disulfide conopeptide with a noncompetitive, potentially allosteric inhibitory mechanism at the nAChRs. The small size and rigidity of the Czon1107 peptide could provide a scaffold for rational drug design strategies for allosteric nAChR modulation. This new paradigm in the "conotoxinomic" structure-function space provides an impetus to screen venom from other Conus species for similar, short bioactive peptides that allosterically modulate ligand-gated receptor function.
- Molecular Biophysics Unit, Indian Institute of Science, Bangalore, Karnataka 560012, India.
Organizational Affiliation: