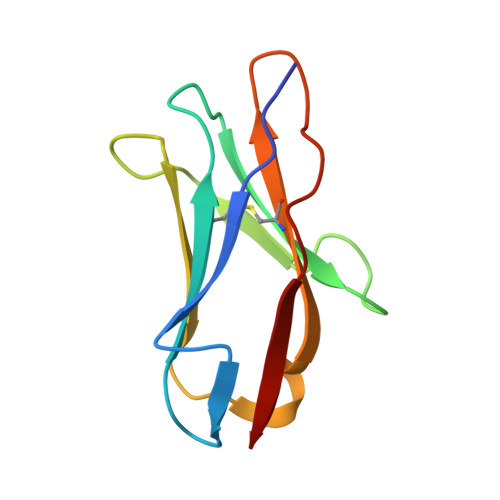
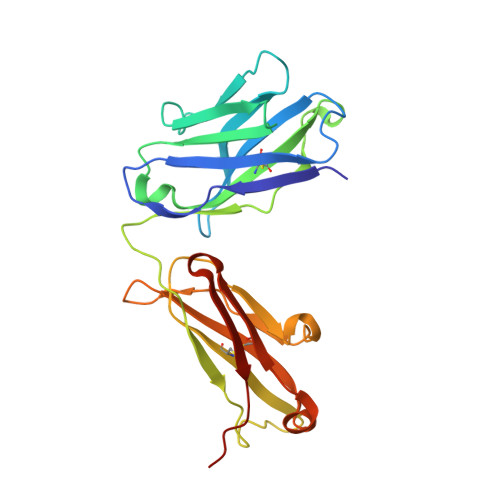
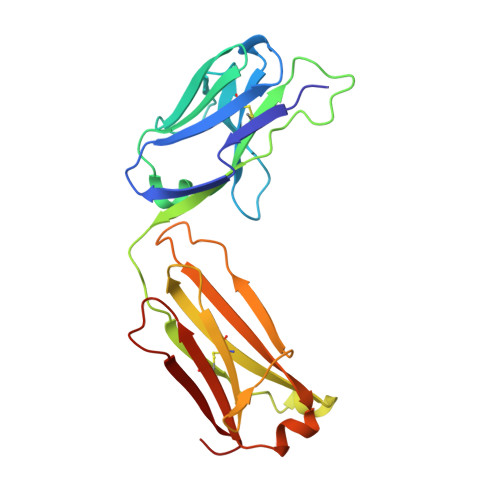
Disrupting LILRB4/APOE Interaction by an Efficacious Humanized Antibody Reverses T-cell Suppression and Blocks AML Development.
Gui, X., Deng, M., Song, H., Chen, Y., Xie, J., Li, Z., He, L., Huang, F., Xu, Y., Anami, Y., Yu, H., Yu, C., Li, L., Yuan, Z., Xu, X., Wang, Q., Chai, Y., Huang, T., Shi, Y., Tsuchikama, K., Liao, X.C., Xia, N., Gao, G.F., Zhang, N., Zhang, C.C., An, Z.(2019) Cancer Immunol Res 7: 1244-1257
- PubMed: 31213474 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1158/2326-6066.CIR-19-0036
- Primary Citation Related Structures:
6K7O - PubMed Abstract:
Therapeutic strategies are urgently needed for patients with acute myeloid leukemia (AML). Leukocyte immunoglobulin-like receptor B4 (LILRB4), which suppresses T-cell activation and supports tissue infiltration of AML cells, represents an attractive drug target for anti-AML therapeutics. Here, we report the identification and development of an LILRB4-specific humanized mAb that blocks LILRB4 activation. This mAb, h128-3, showed potent activity in blocking the development of monocytic AML in various models including patient-derived xenograft mice and syngeneic immunocompetent AML mice. MAb h128-3 enhanced the anti-AML efficacy of chemotherapy treatment by stimulating mobilization of leukemia cells. Mechanistic studies revealed four concordant modes of action for the anti-AML activity of h128-3: (i) reversal of T-cell suppression, (ii) inhibition of monocytic AML cell tissue infiltration, (iii) antibody-dependent cellular cytotoxicity, and (iv) antibody-dependent cellular phagocytosis. Therefore, targeting LILRB4 with antibody represents an effective therapeutic strategy for treating monocytic AML.
- Texas Therapeutics Institute, Brown Foundation Institute of Molecular Medicine, University of Texas Health Science Center, Houston, Texas.
Organizational Affiliation: