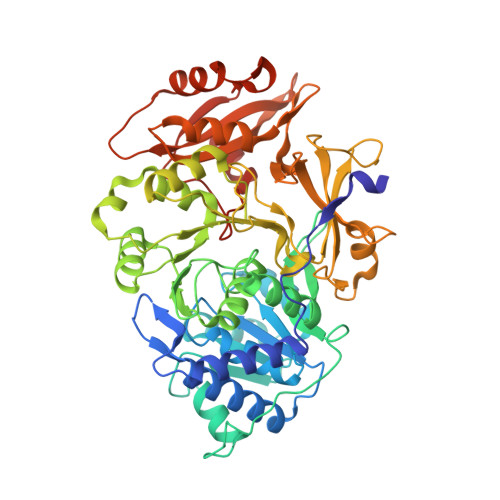
Resurrecting the ancient glow of the fireflies.
Oba, Y., Konishi, K., Yano, D., Shibata, H., Kato, D., Shirai, T.(2020) Sci Adv 6
- PubMed: 33268373
- DOI: https://doi.org/10.1126/sciadv.abc5705
- Primary Citation of Related Structures:
6K4C, 6K4D - PubMed Abstract:
The color of firefly bioluminescence is determined by the structure of luciferase. Firefly luciferase genes have been isolated from more than 30 species, producing light ranging in color from green to orange-yellow. Here, we reconstructed seven ancestral firefly luciferase genes, characterized the enzymatic properties of the recombinant proteins, and determined the crystal structures of the gene from ancestral Lampyridae. Results showed that the synthetic luciferase for the last common firefly ancestor exhibited green light caused by a spatial constraint on the luciferin molecule in enzyme, while fatty acyl-CoA synthetic activity, an original function of firefly luciferase, was diminished in exchange. All known firefly species are bioluminescent in the larvae, with a common ancestor arising approximately 100 million years ago. Combined, our findings propose that, within the mid-Cretaceous forest, the common ancestor of fireflies evolved green light luciferase via trade-off of the original function, which was likely aposematic warning display against nocturnal predation.
- Department of Environmental Biology, Chubu University, Kasugai 487-8501, Japan. yoba@isc.chubu.ac.jp t_shirai@nagahama-i-bio.ac.jp.
Organizational Affiliation: