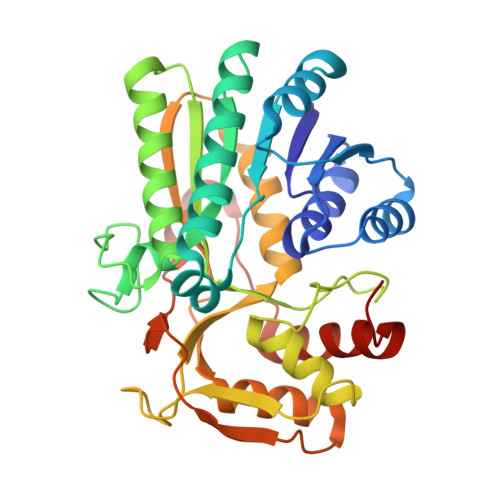
Structural basis for broad substrate specificity of UDP-glucose 4-epimerase in the human milk oligosaccharide catabolic pathway of Bifidobacterium longum.
Nam, Y.W., Nishimoto, M., Arakawa, T., Kitaoka, M., Fushinobu, S.(2019) Sci Rep 9: 11081-11081
- PubMed: 31366978 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-019-47591-w
- Primary Citation Related Structures:
6K0G, 6K0H, 6K0I - PubMed Abstract:
Infant gut-associated bifidobacteria has a metabolic pathway that specifically utilizes lacto-N-biose I (Gal-β1,3-GlcNAc) and galacto-N-biose (Gal-β1,3-GalNAc) from human milk and mucin glycans. UDP-glucose 4-epimerase (GalE) from Bifidobacterium longum (bGalE) catalyzes epimerization reactions of UDP-Gal into UDP-Glc and UDP-GalNAc into UDP-GlcNAc with the same level of activity that is required to send galacto-hexoses into glycolysis. Here, we determined the crystal structures of bGalE in three ternary complex forms: NAD + /UDP, NAD + /UDP-GlcNAc, and NAD + /UDP-Glc. The broad specificity of bGalE was explained by structural features of the binding pocket for the N-acetyl or C2 hydroxy group of the substrate. Asn200 is located in a pocket of the C2 group, and its side chain adopts different conformations in the complex structures with UDP-Glc and UDP-GlcNAc. On the other side, Cys299 forms a large pocket for the C5 sugar ring atom. The flexible C2 pocket and the large C5 pocket of bGalE are suitable for accommodating both the hydroxy and N-acetyl groups of the substrate during sugar ring rotation in the catalytic cycle. The substrate specificity and active site structure of bGalE were distinct from those of Esherichia coli GalE but similar to those of human GalE.
- Department of Biotechnology, The University of Tokyo, Tokyo, 113-8657, Japan.
Organizational Affiliation: