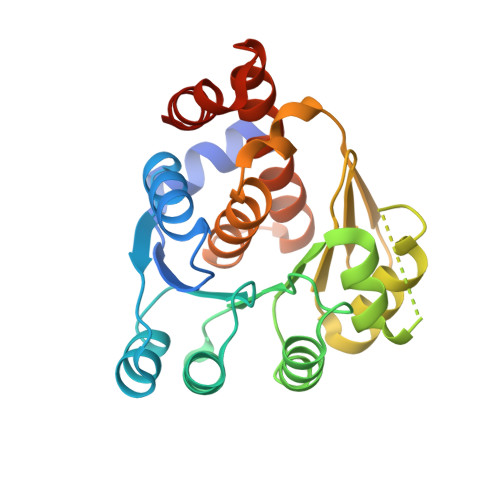
Structural insight of the 5-(Hydroxyethyl)-methylthiazole kinase ThiM involving vitamin B1 biosynthetic pathway from the Klebsiella pneumoniae.
Chen, Y., Wang, L., Shang, F., Liu, W., Lan, J., Chen, J., Ha, N.C., Quan, C., Nam, K.H., Xu, Y.(2019) Biochem Biophys Res Commun 518: 513-518
- PubMed: 31439375 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2019.08.086
- Primary Citation Related Structures:
6JYY, 6K28 - PubMed Abstract:
Thiamin pyrophosphate (TPP) is an essential co-factor in amino acid and carbohydrate metabolic pathways. The TPP-related vitamin B1 biosynthetic pathway is found in most bacterial, plant and lower eukaryotic processes; however, it is not present in humans. In bacterial thiamin synthesis and salvage pathways, the 5-(hydroxyethyl)-methylthiazole kinase (ThiM) is essential in the pathway forming TPP. Thus, ThiM is considered to be an attractive antibacterial drug target. Here, we determined the crystal structures of ThiM from pathogenic Klebsiella pneumoniae (KpThiM) and KpThiM in complex with its substrate 5-(hydroxyethyl)-4-methylthiazole (TZE). KpThiM, consisting of an α-β-α domain, shows a pseudosymmetric trimeric formation. TZE molecules are located in the interface between the KpThiM subunits in the trimer and interact with Met49 and Cys200. Superimposition of the apo and TZE-complexed structures of KpThiM show that the side chains of the amino acids interacting with TZE and Mg 2+ have a rigid configuration. Comparison of the ThiM structures shows that KpThiM could, in terms of sequence and configuration, be different from other ThiM proteins, which possess different amino acids that recognize TZE and Mg 2+ . The structures will provide new insight into the ThiM subfamily proteins for antibacterial drug development.
- Department of Bioengineering, College of Life Science, Dalian Minzu University, Dalian, 116600, Liaoning, China; Key Laboratory of Biotechnology and Bioresources Utilization (Dalian Minzu University), Ministry of Education, China.
Organizational Affiliation: