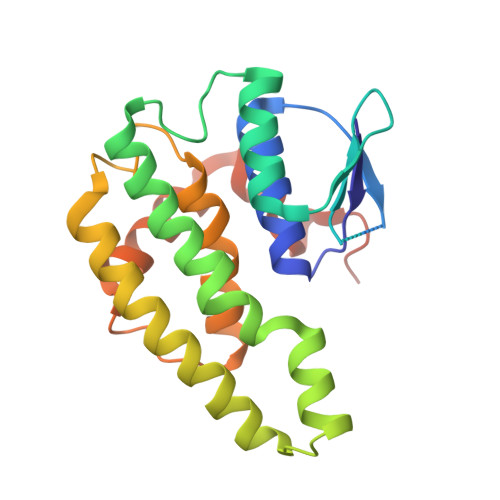
Structure and biochemical studies of a pseudomonad maleylpyruvate isomerase from Pseudomonas aeruginosa PAO1.
Hong, H., Seo, H., Kim, K.J.(2019) Biochem Biophys Res Commun 514: 991-997
- PubMed: 31092332
- DOI: https://doi.org/10.1016/j.bbrc.2019.05.048
- Primary Citation of Related Structures:
6JWK - PubMed Abstract:
Pseudomonas aeruginosa PAO1 can utilize various aromatic hydrocarbons as a carbon source. Among the three genes involved in the gentisate pathway of P. aeruginosa, the gene product of PA2473 belongs to the ζ-class glutathione S-transferase and is predicted to be a maleylpyruvate isomerase. In this study, we determined the crystal structure of maleylpyruvate isomerase from Pseudomonas aeruginosa PAO1 (PaMPI) at a resolution of 1.8 Å. PaMPI functions as a dimer and shows the glutathione S-transferase fold. The structure comparison with other glutathione S-transferase structures enabled us to predict the glutathione cofactor binding site and suggests that PaMPI has differences in residues that make up the putative substrate binding site. Biochemical study of PaMPI showed that the protein has an MPI activity. Interestingly, unlike the reported glutathione S-transferases so far, the purified PaMPI showed isomerase activity without the addition of the reduced glutathione, although the protein showed much higher activity when the glutathione cofactor was added to the reaction mixture. Taken together, our studies reveal that the gene product of PA2473 functions as a maleylpyruvate isomerase and might be involved in the gentisate pathway.
- School of Life Sciences, KNU Creative BioResearch Group, Kyungpook National University, University, Daehak-ro 80, Buk-ku, Daegu, 41566, Republic of Korea; KNU Institute for Microorganisms, Kyungpook National University, Daehak-ro 80, Buk-ku, Daegu, 41566, Republic of Korea.
Organizational Affiliation: