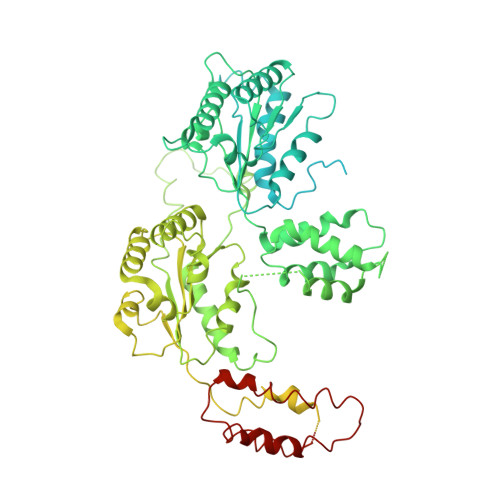
Structural basis of nucleosome assembly by the Abo1 AAA+ ATPase histone chaperone.
Cho, C., Jang, J., Kang, Y., Watanabe, H., Uchihashi, T., Kim, S.J., Kato, K., Lee, J.Y., Song, J.J.(2019) Nat Commun 10: 5764-5764
- PubMed: 31848341 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-019-13743-9
- Primary Citation Related Structures:
6JPQ, 6JPU, 6JQ0 - PubMed Abstract:
The fundamental unit of chromatin, the nucleosome, is an intricate structure that requires histone chaperones for assembly. ATAD2 AAA+ ATPases are a family of histone chaperones that regulate nucleosome density and chromatin dynamics. Here, we demonstrate that the fission yeast ATAD2 homolog, Abo1, deposits histone H3-H4 onto DNA in an ATP-hydrolysis-dependent manner by in vitro reconstitution and single-tethered DNA curtain assays. We present cryo-EM structures of an ATAD2 family ATPase to atomic resolution in three different nucleotide states, revealing unique structural features required for histone loading on DNA, and directly visualize the transitions of Abo1 from an asymmetric spiral (ATP-state) to a symmetric ring (ADP- and apo-states) using high-speed atomic force microscopy (HS-AFM). Furthermore, we find that the acidic pore of ATP-Abo1 binds a peptide substrate which is suggestive of a histone tail. Based on these results, we propose a model whereby Abo1 facilitates H3-H4 loading by utilizing ATP.
- Department of Biological Sciences and KI for the BioCentury, Korea Advanced Institute of Science and Technology (KAIST), Daejeon, Korea. carol.cho@kaist.ac.kr.
Organizational Affiliation: