Crystal structure of GH10 family xylanase XynAF1 from Aspergillus fumigatus Z5
Li, G., Miao, Y., Zhang, R.To be published.
Experimental Data Snapshot
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Beta-xylanase | 319 | Aspergillus fumigatus Z5 | Mutation(s): 0 Gene Names: Y699_04481 EC: 3.2.1.8 | 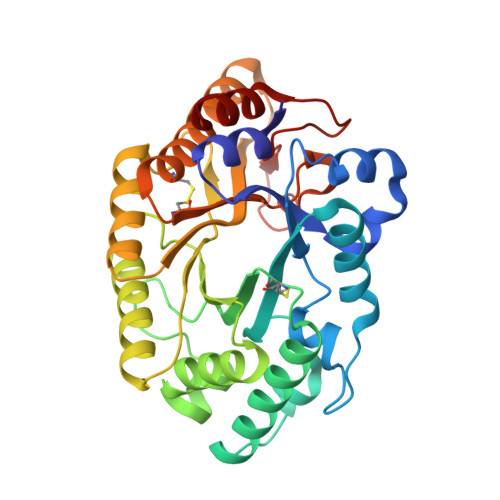 | |
| Ligands 4 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| NAG Download:Ideal Coordinates CCD File | G [auth A], M [auth B] | 2-acetamido-2-deoxy-beta-D-glucopyranose C8 H15 N O6 OVRNDRQMDRJTHS-FMDGEEDCSA-N |  | ||
| MES Download:Ideal Coordinates CCD File | K [auth A] | 2-(N-MORPHOLINO)-ETHANESULFONIC ACID C6 H13 N O4 S SXGZJKUKBWWHRA-UHFFFAOYSA-N |  | ||
| PEG Download:Ideal Coordinates CCD File | L [auth A], P [auth B] | DI(HYDROXYETHYL)ETHER C4 H10 O3 MTHSVFCYNBDYFN-UHFFFAOYSA-N |  | ||
| GOL Download:Ideal Coordinates CCD File | H [auth A], I [auth A], J [auth A], N [auth B], O [auth B] | GLYCEROL C3 H8 O3 PEDCQBHIVMGVHV-UHFFFAOYSA-N |  | ||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| ID | Chains | Name | Type/Class | 2D Diagram | 3D Interactions |
| PRD_900116 Query on PRD_900116 | C, D, E, F | 4beta-beta-xylobiose | Oligosaccharide / Metabolism |  |
| Length ( Å ) | Angle ( ˚ ) |
|---|---|
| a = 44.348 | α = 73.88 |
| b = 57.4 | β = 80.47 |
| c = 64.924 | γ = 68.59 |
| Software Name | Purpose |
|---|---|
| HKL-2000 | data reduction |
| HKL-2000 | data scaling |
| REFMAC | refinement |
| PDB_EXTRACT | data extraction |
| PHASER | phasing |