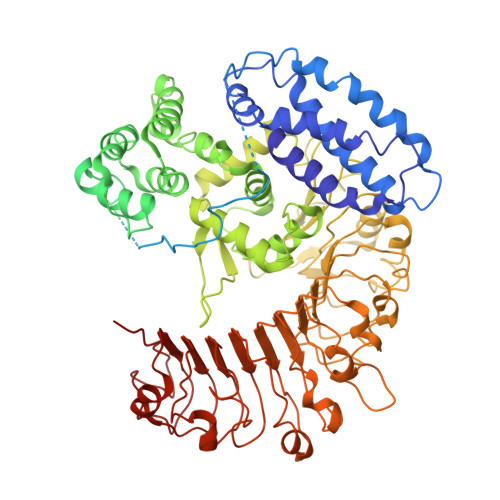
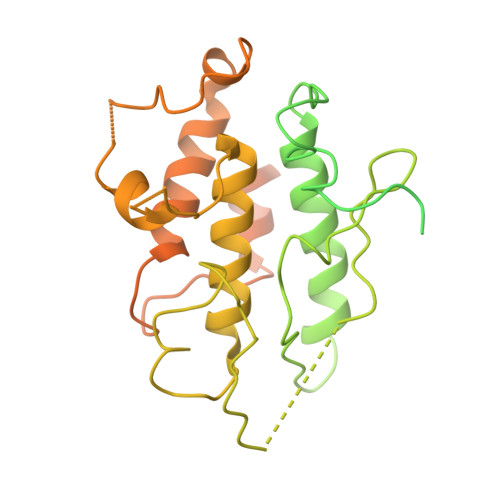
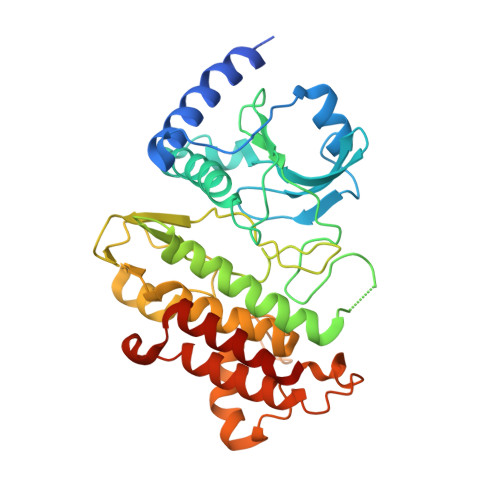
Ligand-triggered allosteric ADP release primes a plant NLR complex.
Wang, J., Wang, J., Hu, M., Wu, S., Qi, J., Wang, G., Han, Z., Qi, Y., Gao, N., Wang, H.W., Zhou, J.M., Chai, J.(2019) Science 364
- PubMed: 30948526 Search on PubMed
- DOI: https://doi.org/10.1126/science.aav5868
- Primary Citation Related Structures:
6J5U, 6J5V, 6J5W - PubMed Abstract:
Pathogen recognition by nucleotide-binding (NB), leucine-rich repeat (LRR) receptors (NLRs) plays roles in plant immunity. The Xanthomonas campestris pv. campestris effector AvrAC uridylylates the Arabidopsis PBL2 kinase, and the latter (PBL2 UMP ) acts as a ligand to activate the NLR ZAR1 precomplexed with the RKS1 pseudokinase. Here we report the cryo-electron microscopy structures of ZAR1-RKS1 and ZAR1-RKS1-PBL2 UMP in an inactive and intermediate state, respectively. The ZAR1 LRR domain, compared with animal NLR LRR domains, is differently positioned to sequester ZAR1 in an inactive state. Recognition of PBL2 UMP is exclusively through RKS1, which interacts with ZAR1 LRR PBL2 UMP binding stabilizes the RKS1 activation segment, which sterically blocks ZAR1 adenosine diphosphate (ADP) binding. This engenders a more flexible NB domain without conformational changes in the other ZAR1 domains. Our study provides a structural template for understanding plant NLRs.
- State Key Laboratory of Plant Genomics, Institute of Genetics and Developmental Biology, Academy of Seed Design, Chinese Academy of Sciences, 100101 Beijing, China.
Organizational Affiliation: